| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Gastrin/cholecystokinin type B receptor |
|---|
| Ligand | BDBM50048803 |
|---|
| Substrate/Competitor | n/a |
|---|
| Ki | 1000±n/a nM |
|---|
| Comments | PDSP_1996 |
|---|
| Citation |  Schotte, A; Janssen, PF; Gommeren, W; Luyten, WH; Van Gompel, P; Lesage, AS; De Loore, K; Leysen, JE Risperidone compared with new and reference antipsychotic drugs: in vitro and in vivo receptor binding. Psychopharmacology (Berl)124:57-73 (1996) [PubMed] Article Schotte, A; Janssen, PF; Gommeren, W; Luyten, WH; Van Gompel, P; Lesage, AS; De Loore, K; Leysen, JE Risperidone compared with new and reference antipsychotic drugs: in vitro and in vivo receptor binding. Psychopharmacology (Berl)124:57-73 (1996) [PubMed] Article |
|---|
| More Info.: | Get all data from this article |
|---|
| |
| Gastrin/cholecystokinin type B receptor |
|---|
| Name: | Gastrin/cholecystokinin type B receptor |
|---|
| Synonyms: | Cckbr | Cholecystokinin A | Cholecystokinin B receptor | Cholecystokinin receptor | GASR_RAT | Gastrin/cholecystokinin type B receptor |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 48980.43 |
|---|
| Organism: | RAT |
|---|
| Description: | Cholecystokinin A CCKBR RAT::P30553 |
|---|
| Residue: | 452 |
|---|
| Sequence: | MELLKLNRSVQGPGPGSGSSLCRPGVSLLNSSSAGNLSCDPPRIRGTGTRELEMAIRITL
YAVIFLMSVGGNVLIIVVLGLSRRLRTVTNAFLLSLAVSDLLLAVACMPFTLLPNLMGTF
IFGTVICKAISYLMGVSVSVSTLNLVAIALERYSAICRPLQARVWQTRSHAARVILATWL
LSGLLMVPYPVYTMVQPVGPRVLQCMHRWPSARVQQTWSVLLLLLLFFIPGVVIAVAYGL
ISRELYLGLHFDGENDSETQSRARNQGGLPGGAAPGPVHQNGGCRPVTSVAGEDSDGCCV
QLPRSRLEMTTLTTPTPGPVPGPRPNQAKLLAKKRVVRMLLVIVLLFFLCWLPVYSVNTW
RAFDGPGAQRALSGAPISFIHLLSYVSACVNPLVYCFMHRRFRQACLDTCARCCPRPPRA
RPQPLPDEDPPTPSIASLSRLSYTTISTLGPG
|
|
|
|---|
| BDBM50048803 |
|---|
| n/a |
|---|
| Name | BDBM50048803 |
|---|
| Synonyms: | 5-(2-(4-(benzo[d]isothiazol-3-yl)piperazin-1-yl)ethyl)-6-chloroindolin-2-one | 5-[2-(4-Benzo[d]isothiazol-3-yl-piperazin-1-yl)-ethyl]-6-chloro-1,3-dihydro-indol-2-one | 5-[2-(4-Benzo[d]isothiazol-3-yl-piperazin-1-yl)-ethyl]-6-chloro-1,3-dihydro-indol-2-one (Ziprasidone) | 5-[2-(4-Benzo[d]isothiazol-3-yl-piperazin-1-yl)-ethyl]-6-chloro-1,3-dihydro-indol-2-one(Norastemizole) | CHEMBL708 | GEODON | ZIPRASIDONE | ZIPRASIDONE HYDROCHLORIDE |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H21ClN4OS |
|---|
| Mol. Mass. | 412.936 |
|---|
| SMILES | Clc1cc2NC(=O)Cc2cc1CCN1CCN(CC1)c1nsc2ccccc12 |
|---|
| Structure | 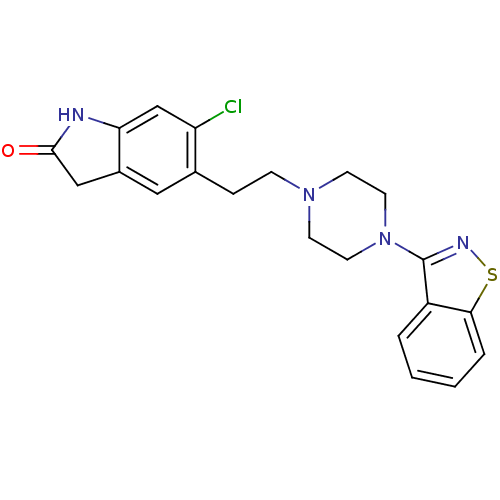
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Schotte, A; Janssen, PF; Gommeren, W; Luyten, WH; Van Gompel, P; Lesage, AS; De Loore, K; Leysen, JE Risperidone compared with new and reference antipsychotic drugs: in vitro and in vivo receptor binding. Psychopharmacology (Berl)124:57-73 (1996) [PubMed] Article
Schotte, A; Janssen, PF; Gommeren, W; Luyten, WH; Van Gompel, P; Lesage, AS; De Loore, K; Leysen, JE Risperidone compared with new and reference antipsychotic drugs: in vitro and in vivo receptor binding. Psychopharmacology (Berl)124:57-73 (1996) [PubMed] Article