| Reaction Details |
|---|
 | Report a problem with these data |
| Target | ADP-ribosylation factor GTPase-activating protein 1 |
|---|
| Ligand | BDBM95229 |
|---|
| Substrate/Competitor | n/a |
|---|
| IC50 | >18306±n/a nM |
|---|
| Citation |  PubChem, PC Fluorescence polarization-based biochemical high throughput dose response screening assay to identify inhibitors of ADP-ribosylation factor GTPase activating protein 1 (ARFGAP1) PubChem Bioassay(2012)[AID] PubChem, PC Fluorescence polarization-based biochemical high throughput dose response screening assay to identify inhibitors of ADP-ribosylation factor GTPase activating protein 1 (ARFGAP1) PubChem Bioassay(2012)[AID] |
|---|
| More Info.: | Get all data from this article |
|---|
| |
| ADP-ribosylation factor GTPase-activating protein 1 |
|---|
| Name: | ADP-ribosylation factor GTPase-activating protein 1 |
|---|
| Synonyms: | ARFG1_RAT | Arf1gap | Arfgap1 |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 45436.38 |
|---|
| Organism: | Rattus norvegicus |
|---|
| Description: | gi_21489979 |
|---|
| Residue: | 415 |
|---|
| Sequence: | MASPRTRKVLKEVRAQDENNVCFECGAFNPQWVSVTYGIWICLECSGRHRGLGVHLSFVR
SVTMDKWKDIELEKMKAGGNAKFREFLEAQDDYEPSWSLQDKYSSRAAALFRDKVATLAE
GKEWSLESSPAQNWTPPQPKTLQFTAHRPAGQPQNVTTSGDKAFEDWLNDDLGSYQGAQE
NRYVGFGNTVPPQKREDDFLNSAMSSLYSGWSSFTTGASKFASAAKEGATKFGSQASQKA
SELGHSLNENVLKPAQEKVKEGRIFDDVSSGVSQLASKVQGVGSKGWRDVTTFFSGKAED
TSDRPLEGHSYQNSSGDNSQNSTIDQSFWETFGSAEPPKAKSPSSDSWTCADASTGRRSS
DSWDIWGSGSASNNKNSNSDGWESWEGASGEGRAKATKKAAPSTAADEGWDNQNW
|
|
|
|---|
| BDBM95229 |
|---|
| n/a |
|---|
| Name | BDBM95229 |
|---|
| Synonyms: | MLS001234985 | N-(3-chlorophenyl)-2-(7-methyl-4-oxidanylidene-pyrido[1,2-a][1,3,5]triazin-2-yl)sulfanyl-ethanamide | N-(3-chlorophenyl)-2-(7-methyl-4-oxopyrido[1,2-a][1,3,5]triazin-2-yl)sulfanylacetamide | N-(3-chlorophenyl)-2-[(4-keto-7-methyl-pyrido[1,2-a][1,3,5]triazin-2-yl)thio]acetamide | N-(3-chlorophenyl)-2-[(7-methyl-4-oxo-2-pyrido[1,2-a][1,3,5]triazinyl)thio]acetamide | SMR000810925 | cid_7259073 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C16H13ClN4O2S |
|---|
| Mol. Mass. | 360.818 |
|---|
| SMILES | Cc1ccc2nc(SCC(=O)Nc3cccc(Cl)c3)nc(=O)n2c1 |
|---|
| Structure | 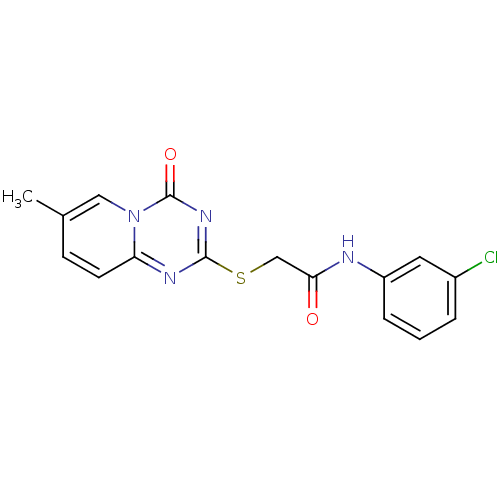
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Fluorescence polarization-based biochemical high throughput dose response screening assay to identify inhibitors of ADP-ribosylation factor GTPase activating protein 1 (ARFGAP1) PubChem Bioassay(2012)[AID]
PubChem, PC Fluorescence polarization-based biochemical high throughput dose response screening assay to identify inhibitors of ADP-ribosylation factor GTPase activating protein 1 (ARFGAP1) PubChem Bioassay(2012)[AID]