| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Nuclear receptor subfamily 5 group A member 2 |
|---|
| Ligand | BDBM95529 |
|---|
| Substrate/Competitor | n/a |
|---|
| IC50 | 1374±n/a nM |
|---|
| Citation |  PubChem, PC Luminescence-based cell-based high throughput dose response assay for inverse agonists of the liver receptor homolog-1 (LRH-1; NR5A2) PubChem Bioassay(2013)[AID] PubChem, PC Luminescence-based cell-based high throughput dose response assay for inverse agonists of the liver receptor homolog-1 (LRH-1; NR5A2) PubChem Bioassay(2013)[AID] |
|---|
| More Info.: | Get all data from this article |
|---|
| |
| Nuclear receptor subfamily 5 group A member 2 |
|---|
| Name: | Nuclear receptor subfamily 5 group A member 2 |
|---|
| Synonyms: | Alpha-1-fetoprotein transcription factor | B1-binding factor | B1F | CPF | CYP7A promoter-binding factor | FTF | Hepatocytic transcription factor | LRH-1 | Liver Receptor Homolog 1 | NR5A2 | NR5A2_HUMAN | Nuclear receptor subfamily 5 group A member 2 | hB1F | nuclear receptor subfamily 5 group A member 2 isoform 2 |
|---|
| Type: | Nuclear Hormone Receptor |
|---|
| Mol. Mass.: | 61341.37 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 541 |
|---|
| Sequence: | MSSNSDTGDLQESLKHGLTPIGAGLPDRHGSPIPARGRLVMLPKVETEALGLARSHGEQG
QMPENMQVSQFKMVNYSYDEDLEELCPVCGDKVSGYHYGLLTCESCKGFFKRTVQNNKRY
TCIENQNCQIDKTQRKRCPYCRFQKCLSVGMKLEAVRADRMRGGRNKFGPMYKRDRALKQ
QKKALIRANGLKLEAMSQVIQAMPSDLTISSAIQNIHSASKGLPLNHAALPPTDYDRSPF
VTSPISMTMPPHGSLQGYQTYGHFPSRAIKSEYPDPYTSSPESIMGYSYMDSYQTSSPAS
IPHLILELLKCEPDEPQVQAKIMAYLQQEQANRSKHEKLSTFGLMCKMADQTLFSIVEWA
RSSIFFRELKVDDQMKLLQNCWSELLILDHIYRQVVHGKEGSIFLVTGQQVDYSIIASQA
GATLNNLMSHAQELVAKLRSLQFDQREFVCLKFLVLFSLDVKNLENFQLVEGVQEQVNAA
LLDYTMCNYPQQTEKFGQLLLRLPEIRAISMQAEEYLYYKHLNGDVPYNNLLIEMLHAKR
A
|
|
|
|---|
| BDBM95529 |
|---|
| n/a |
|---|
| Name | BDBM95529 |
|---|
| Synonyms: | MLS001018993 | N-(4-methoxyphenyl)-2-(1H-pyrazolo[3,4-d]pyrimidin-4-ylsulfanyl)acetamide | N-(4-methoxyphenyl)-2-(1H-pyrazolo[3,4-d]pyrimidin-4-ylsulfanyl)ethanamide | N-(4-methoxyphenyl)-2-(1H-pyrazolo[3,4-d]pyrimidin-4-ylthio)acetamide | SMR000363374 | cid_2475269 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C14H13N5O2S |
|---|
| Mol. Mass. | 315.35 |
|---|
| SMILES | COc1ccc(NC(=O)CSc2ncnc3n[nH]cc23)cc1 |
|---|
| Structure | 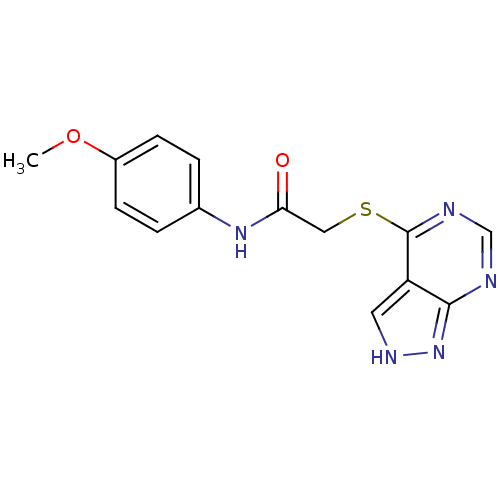
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Luminescence-based cell-based high throughput dose response assay for inverse agonists of the liver receptor homolog-1 (LRH-1; NR5A2) PubChem Bioassay(2013)[AID]
PubChem, PC Luminescence-based cell-based high throughput dose response assay for inverse agonists of the liver receptor homolog-1 (LRH-1; NR5A2) PubChem Bioassay(2013)[AID]