| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Metabotropic glutamate receptor 2 |
|---|
| Ligand | BDBM160721 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | MGluR2 cAMP Assay |
|---|
| pH | 7±n/a |
|---|
| EC50 | 280±n/a nM |
|---|
| Comments | extracted |
|---|
| Citation |  Mattson, RJ; Meng, Z; Guernon, LR Positive allosteric modulators of mGluR2 US Patent US9682980 Publication Date 6/20/2017 Mattson, RJ; Meng, Z; Guernon, LR Positive allosteric modulators of mGluR2 US Patent US9682980 Publication Date 6/20/2017 |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Metabotropic glutamate receptor 2 |
|---|
| Name: | Metabotropic glutamate receptor 2 |
|---|
| Synonyms: | GPRC1B | GRM2 | GRM2_HUMAN | MGLUR2 | Metabotropic glutamate receptor | glutamate receptor, metabotropic 2 precursor | mGlu2 | metabotropic glutamate 2 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 95584.88 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q14416 |
|---|
| Residue: | 872 |
|---|
| Sequence: | MGSLLALLALLLLWGAVAEGPAKKVLTLEGDLVLGGLFPVHQKGGPAEDCGPVNEHRGIQ
RLEAMLFALDRINRDPHLLPGVRLGAHILDSCSKDTHALEQALDFVRASLSRGADGSRHI
CPDGSYATHGDAPTAITGVIGGSYSDVSIQVANLLRLFQIPQISYASTSAKLSDKSRYDY
FARTVPPDFFQAKAMAEILRFFNWTYVSTVASEGDYGETGIEAFELEARARNICVATSEK
VGRAMSRAAFEGVVRALLQKPSARVAVLFTRSEDARELLAASQRLNASFTWVASDGWGAL
ESVVAGSEGAAEGAITIELASYPISDFASYFQSLDPWNNSRNPWFREFWEQRFRCSFRQR
DCAAHSLRAVPFEQESKIMFVVNAVYAMAHALHNMHRALCPNTTRLCDAMRPVNGRRLYK
DFVLNVKFDAPFRPADTHNEVRFDRFGDGIGRYNIFTYLRAGSGRYRYQKVGYWAEGLTL
DTSLIPWASPSAGPLPASRCSEPCLQNEVKSVQPGEVCCWLCIPCQPYEYRLDEFTCADC
GLGYWPNASLTGCFELPQEYIRWGDAWAVGPVTIACLGALATLFVLGVFVRHNATPVVKA
SGRELCYILLGGVFLCYCMTFIFIAKPSTAVCTLRRLGLGTAFSVCYSALLTKTNRIARI
FGGAREGAQRPRFISPASQVAICLALISGQLLIVVAWLVVEAPGTGKETAPERREVVTLR
CNHRDASMLGSLAYNVLLIALCTLYAFKTRKCPENFNEAKFIGFTMYTTCIIWLAFLPIF
YVTSSDYRVQTTTMCVSVSLSGSVVLGCLFAPKLHIILFQPQKNVVSHRAPTSRFGSAAA
RASSSLGQGSGSQFVPTVCNGREVVDSTTSSL
|
|
|
|---|
| BDBM160721 |
|---|
| n/a |
|---|
| Name | BDBM160721 |
|---|
| Synonyms: | US9682980, 8-Chloro-3-(cyclopropylmethyl)-7-(4-(pyrimidin-2-yl)piperazin-1-yl)-[1,2,4]triazolo[4,3-b]pyridazine |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C18H20ClN7 |
|---|
| Mol. Mass. | 369.851 |
|---|
| SMILES | Clc1c(cnn2c(CC3CC3)nnc12)N1CCC(CC1)c1ncccn1 |
|---|
| Structure | 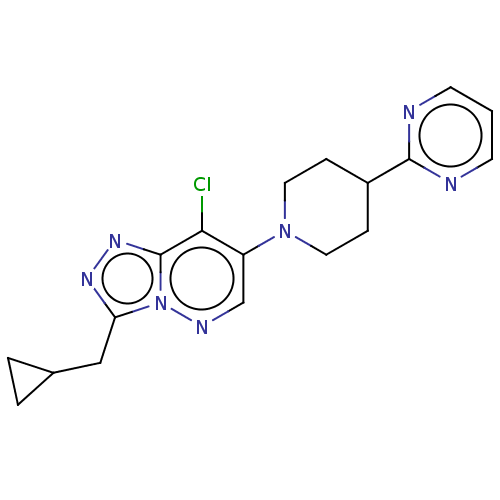
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Mattson, RJ; Meng, Z; Guernon, LR Positive allosteric modulators of mGluR2 US Patent US9682980 Publication Date 6/20/2017
Mattson, RJ; Meng, Z; Guernon, LR Positive allosteric modulators of mGluR2 US Patent US9682980 Publication Date 6/20/2017