null
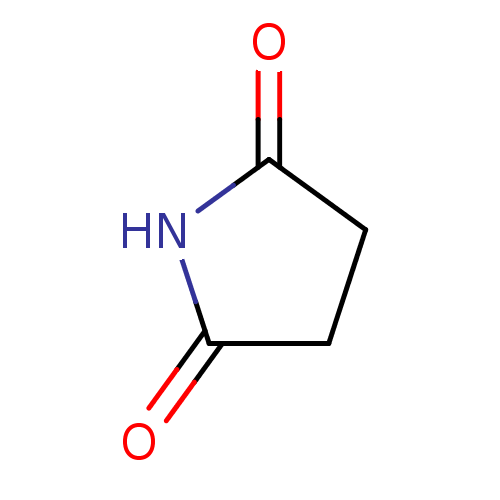
SMILES O=C1CCC(=O)N1
InChI Key InChIKey=KZNICNPSHKQLFF-UHFFFAOYSA-N
PDB links: 20 PDB IDs contain this monomer as substructures. 20 PDB IDs contain inhibitors having a similarity of 90% to this monomer.
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 5 hits for monomerid = 7814
Found 5 hits for monomerid = 7814
Affinity DataKi: 642nMAssay Description:Binding affinity to Magnetospirillum gryphiswaldense cereblon isoform 4 by measuring baseline corrected normalized fluorescence by MST based assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.30E+3nMAssay Description:Inhibition of MANT-uracil binding to wild-type Magnetospirillum gryphiswaldense CRBN isoform 4 by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.30E+3nMAssay Description:Inhibition of MANT-uracil binding to wild type Magnetospirillum gryphiswaldense cereblon isoform 4 by Cheng-Prusoff equation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 7.60E+3nMAssay Description:Inhibition of MANT-uracil binding to wild type Magnetospirillum gryphiswaldense cereblon isoform 4 by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.53E+3nMAssay Description:Binding affinity to Magnetospirillum gryphiswaldense cereblon isoform 4 by measuring baseline corrected normalized fluorescence by MST based assayMore data for this Ligand-Target Pair