null
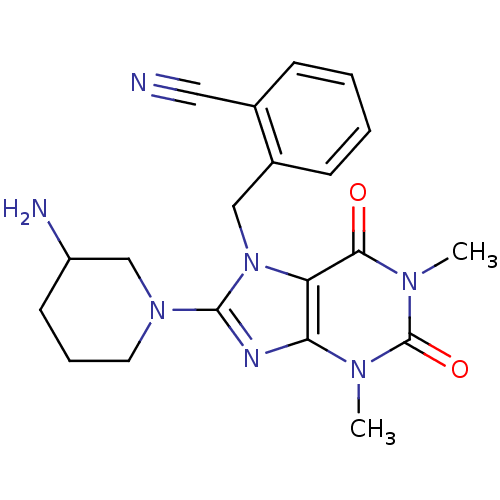
SMILES Cn1c2nc(N3CCCC(N)C3)n(Cc3ccccc3C#N)c2c(=O)n(C)c1=O
InChI Key InChIKey=XJNKUWDMCBZMTG-UHFFFAOYSA-N
PDB links: 1 PDB ID matches this monomer.
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 6 hits for monomerid = 11645
Found 6 hits for monomerid = 11645
Affinity DataKi: 2nM ΔG°: -11.7kcal/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -11.7kcal/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -11.5kcal/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -11.5kcal/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-6.10kcal/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-6.10kcal/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)