null
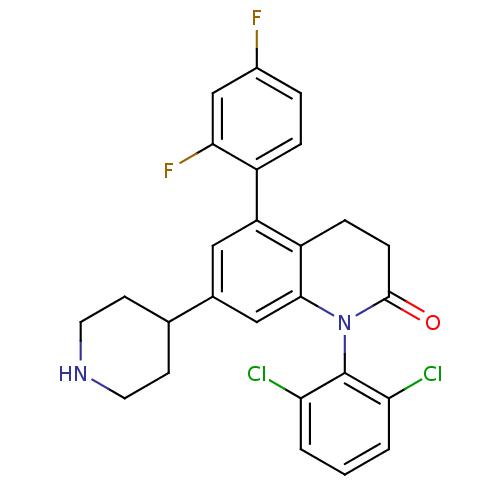
SMILES Fc1ccc(c(F)c1)-c1cc(cc2N(C(=O)CCc12)c1c(Cl)cccc1Cl)C1CCNCC1
InChI Key InChIKey=VXIYTVJEIXMAQF-UHFFFAOYSA-N
PDB links: 1 PDB ID matches this monomer.
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 4 hits for monomerid = 15242
Found 4 hits for monomerid = 15242
Affinity DataIC50: 0.740nMpH: 7.0 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with wild-type or mutant p38alpha enzymes, and substrates in the presence ATP/[gam...More data for this Ligand-Target Pair
Affinity DataIC50: 0.740nMAssay Description:Inhibition of p38alpha MAP kinaseMore data for this Ligand-Target Pair
Target InfoPDBMMDBNCI pathwayReactome pathwayKEGG
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
In DepthDetails
Article
BindingDB Entry DOI: 10.7270/Q2MW2HKRPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)
BindingDB Entry DOI: 10.7270/Q2MW2HKRPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)Affinity DataIC50: 14nMpH: 7.0 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with wild-type or mutant p38alpha enzymes, and substrates in the presence ATP/[gam...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40nMpH: 7.0 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with wild-type or mutant p38alpha enzymes, and substrates in the presence ATP/[gam...More data for this Ligand-Target Pair