null
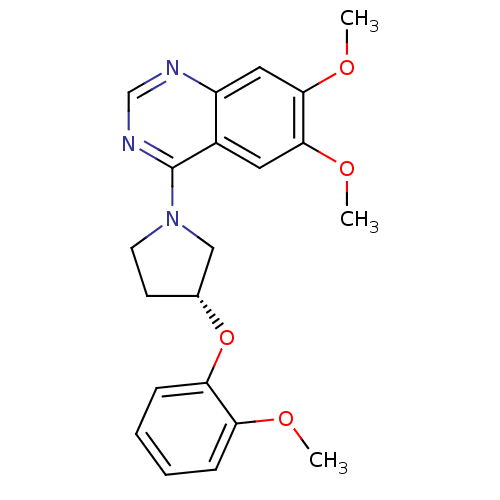
SMILES COc1ccccc1O[C@@H]1CCN(C1)c1ncnc2cc(OC)c(OC)cc12
InChI Key InChIKey=APOQAWXVAQVYAS-CQSZACIVSA-N
PDB links: 1 PDB ID contains inhibitors having a similarity of 90% to this monomer.
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 5 hits for monomerid = 14741
Found 5 hits for monomerid = 14741
Affinity DataKi: >100nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
Affinity DataKi: <100nMAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Homo sapiens (Human))
St. John's University
Curated by ChEMBL
St. John's University
Curated by ChEMBL
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Pfizer
Pfizer
Affinity DataKi: 240nM ΔG°: -8.93kcal/molepH: 7.5 T: 2°CAssay Description:PDE activity was monitored by measuring the hydrolysis of [3H]-cAMP to [3H]-AMP using a scintillation proximity assay (SPA). [3H]-AMP was captured by...More data for this Ligand-Target Pair