null
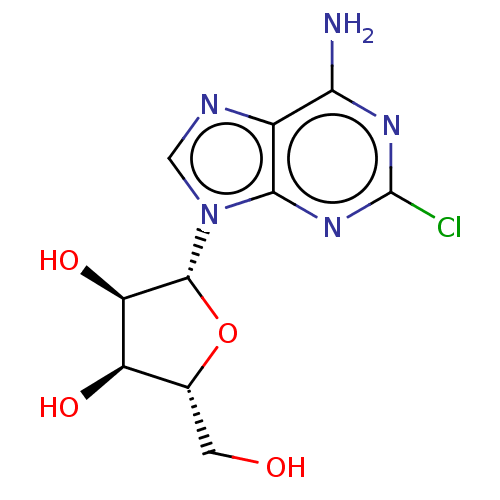
SMILES Nc1nc(Cl)nc2n(cnc12)[C@@H]1O[C@H](CO)[C@@H](O)[C@H]1O
InChI Key InChIKey=BIXYYZIIJIXVFW-UUOKFMHZSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 51 hits for monomerid = 50009525
Found 51 hits for monomerid = 50009525
Affinity DataKi: 1.30nMAssay Description:Antagonist binding of 2-chloro-[3H]-adenosine in rat brainMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Binding affinity to human recombinant adenosine A1 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Displacement of [3H]N6-phenylisopropyladenosine from recombinant human A1AR expressed in CHO cell membranes after 60 mins by liquid scintillation cou...More data for this Ligand-Target Pair
Affinity DataKi: 6.70nMAssay Description:Inhibition of [3H]- (R)-P1A binding to adenosine A1 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Antagonist binding of N6-cyclohexyl-[3H]-adenosine to guinea pig brainMore data for this Ligand-Target Pair
Affinity DataKi: 8.70nMAssay Description:Displacement of [3H]CHA from adenosine A1 receptor of rat whole brainMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Antagonist binding of L-N6-phenyl-isopropyl)-[3H]adenosine to rat fatMore data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Inhibition of [3H]PIA binding to rat cortical adenosine A1 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Binding affinity to adenosine A1 receptor in rat brain membranes by measuring displacement of specific [3H]PIA as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Tested for the binding affinity of A1 receptor by displacing the [3H]-PIA in rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Inhibition of binding of [3H]N6-cyclohexyladenosine to adenosine A1 receptor of rat whole brain membranes.Checked by AuthorMore data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Binding affinity against adenosine A1 receptor using [3H]-CHA or [3H]PIA as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Binding affinity towards adenosine A1 receptor on rat whole brain membrane using [3H]N6-cyclohexyladenosineMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Antagonist binding of 2-chloro-[3H]-adenosine to rat brainMore data for this Ligand-Target Pair
Affinity DataKi: 19nMAssay Description:Binding affinity to human recombinant adenosine A3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Inhibitory activity at Adenosine A1 receptor by inhibition of [3H]CHA binding to bovine brain cortical membranes.More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Antagonist binding of N6-cyclohexyl-[3H]-adenosine to bovine brainMore data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Displacement of [3H]-NECA from adenosine A2 receptor of rat striatal membraneMore data for this Ligand-Target Pair
TargetAdenosine receptor A2a(Rattus norvegicus (rat))
National Institute of Diabetes
Curated by ChEMBL
National Institute of Diabetes
Curated by ChEMBL
Affinity DataKi: 63nMAssay Description:Tested for the binding affinity of A2a receptor by displacing the [3H]-CGS- 21680 in rat striatal membranesMore data for this Ligand-Target Pair
TargetAdenosine receptor A2a(Rattus norvegicus (rat))
National Institute of Diabetes
Curated by ChEMBL
National Institute of Diabetes
Curated by ChEMBL
Affinity DataKi: 63nMAssay Description:Binding affinity to adenosine A2A receptor in rat striatal membranes by measuring displacement of specific [3H]-CGS- 21680 as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Binding affinity towards adenosine A2 receptor on rat striatal membrane using [3H]NECA as radioligandMore data for this Ligand-Target Pair
TargetAdenosine receptor A2a(Rattus norvegicus (rat))
National Institute of Diabetes
Curated by ChEMBL
National Institute of Diabetes
Curated by ChEMBL
TargetAdenosine receptor A2a(Rattus norvegicus (rat))
National Institute of Diabetes
Curated by ChEMBL
National Institute of Diabetes
Curated by ChEMBL
TargetAdenosine receptor A2a(Rattus norvegicus (rat))
National Institute of Diabetes
Curated by ChEMBL
National Institute of Diabetes
Curated by ChEMBL
Affinity DataKi: 76nMAssay Description:Inhibition of [3H]- NECA binding to adenosine receptor A2AMore data for this Ligand-Target Pair
Affinity DataKi: 87nMAssay Description:Affinity for human Adenosine A3 receptor expressed in CHO cellMore data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Inhibitory activity at Adenosine A2A receptor by inhibition of [3H]-CGS- 21680 binding to bovine striatal membranes.More data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Binding affinity to human recombinant adenosine receptor A2aMore data for this Ligand-Target Pair
Affinity DataKi: 650nMAssay Description:Binding affinity at wild-type Adenosine A3 receptor expressed in COS-7 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 920nMAssay Description:Antagonist binding of L-N6-phenyl-isopropyl)-[3H]adenosine to rat brainMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:Displacement of FITC-geldanamycin from GRP94 (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.89E+3nMAssay Description:Tested for the binding affinity of A3 receptor by displacing N6-[[125I]-4-amino-3-iodobenzyl]-adenosine-5''-N-methyluronamide from membranes of CHO ...More data for this Ligand-Target Pair
Affinity DataKi: 1.89E+3nMAssay Description:Binding affinity determined by displacement of specific binding of [125I]N-(4-amino-3-iodophenethyl)-adenosine in membranes of CHO cells stably trans...More data for this Ligand-Target Pair
Affinity DataKi: 2.20E+3nMAssay Description:Displacement of FITC-geldanamycin from HSP90alpha (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.20E+3nMAssay Description:Inhibitory activity against Adenosine A3 receptor by inhibiting specific [3H](R)-PIA binding to rat testis membranesMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 2A(Homo sapiens (Human))
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]ketanserin from human 5-HT2A receptor expressed in HEK-T cell membranes incubated for 90 mins under dark condition by microbeta s...More data for this Ligand-Target Pair
Affinity DataKi: 1.16E+4nMAssay Description:Binding affinity at Mutant (H272E) human adenosine A3 receptor expressed in COS-7 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.14E+4nMAssay Description:Displacement of [3H]PSB-603 from human recombinant adenosine A2B receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.55E+4nMAssay Description:Displacement of [125I]I-ABOPX from human recombinant adenosine A2B receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.45E+4nMAssay Description:Inhibition of calf placental adenosine deaminase by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
Affinity DataKd: 3.80E+4nMAssay Description:Displacement of FITC-geldanamycin from HSP90alpha (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 8.60E+3nMAssay Description:Displacement of FITC-geldanamycin from GRP94 (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetcGMP-dependent 3',5'-cyclic phosphodiesterase(Homo sapiens (Human))
East China University of Science and Technology
Curated by ChEMBL
East China University of Science and Technology
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of recombinant human PDE2A catalytic domain (580 to 919 residues) expressed in Escherichia coli BL21 (Codonplus) using [3H]cGMP as substra...More data for this Ligand-Target Pair
Affinity DataEC50: 1.89E+5nMAssay Description:Agonist activity at human adenosine A2B receptor expressed in CHO cells assessed as stimulation of adenylate cyclaseMore data for this Ligand-Target Pair
Affinity DataEC50: 8.00E+3nMAssay Description:Antagonism of cyclic [3H]AMP accumulation elicited by 15 uM 2-chloroadenosine in [3H]adenine-labeled guinea pig cerebral cortical slices at A2 recept...More data for this Ligand-Target Pair
Affinity DataEC50: 191nMAssay Description:Prolongation of the stimulus-QRS interval by 50% of the maximum response at the adenosine A1 receptor in langendorff guinea pig heart preparationChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Concentration required for 50% inhibition of [3H]-CHA binding on rat brain adenosine A1 receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 1.63E+3nMAssay Description:Effect on forskolin-stimulated cyclic AMP production in intact chinese hamster ovary (CHO) cell expressing the human Adenosine A3 receptorMore data for this Ligand-Target Pair