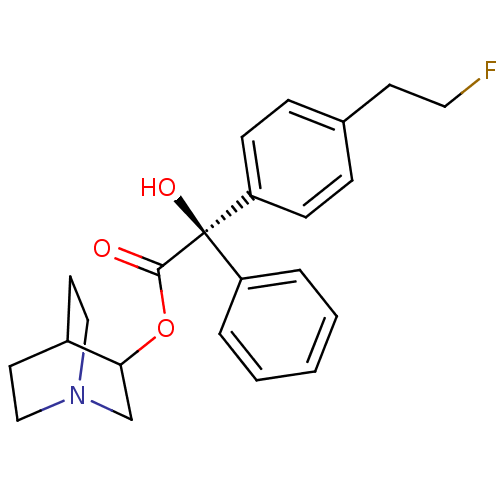
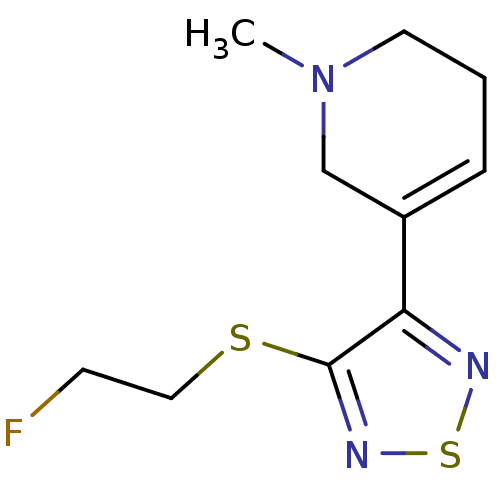
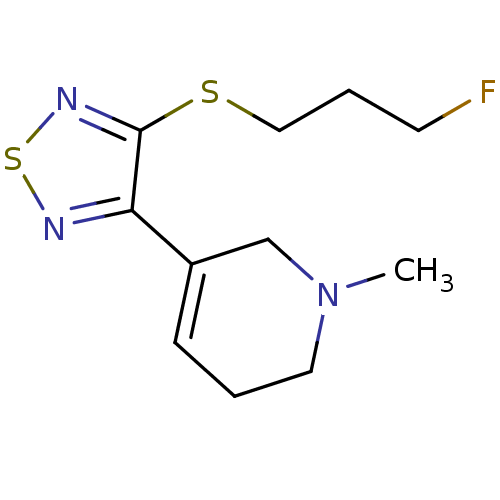
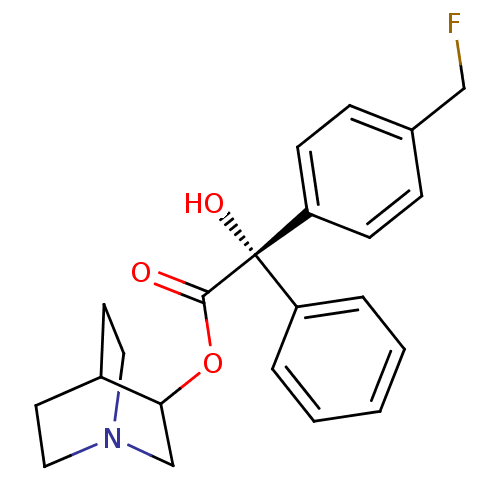
Affinity DataKi: 0.180nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 0.220nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 0.310nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 0.340nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 0.380nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 0.840nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M2 was determined by measuring its ability to displace [3H]-AF-DX 384 from rat he...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M2 was determined by measuring its ability to displace [3H]-AF-DX 384 from rat he...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M2 was determined by measuring its ability to displace [3H]-AF-DX 384 from rat he...More data for this Ligand-Target Pair
Affinity DataKi: 2.30nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M1 was determined by measuring its ability to displace [3H]-Pirenzepine from bovi...More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M2 was determined by measuring its ability to displace [3H]-AF-DX 384 from rat he...More data for this Ligand-Target Pair
Affinity DataKi: 4.20nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 4.60nMAssay Description:In vitro binding affinity against against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 5.5nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 5.5nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 6.30nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M2 was determined by measuring its ability to displace [3H]-AF-DX 384 from rat he...More data for this Ligand-Target Pair
Affinity DataKi: 7.40nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M1 was determined by measuring its ability to displace [3H]-Pirenzepine from bovi...More data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 8.10nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:In vitro binding affinity against rat heart membrane using [3H]-AF-DX-384More data for this Ligand-Target Pair
Affinity DataKi: 17.9nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
Affinity DataKi: 17.9nMAssay Description:In vitro binding affinity against bovine striatal membrane using [3H]-pirenzepineMore data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Homo sapiens (Human))
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 18.6nMAssay Description:In vitro binding affinity towards Sigma receptor type 1 was determined by measuring its ability to displace [3H]-(+)-pentazocine from guinea pig brai...More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Homo sapiens (Human))
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 21.7nMAssay Description:In vitro binding affinity towards Sigma receptor type 1 was determined by measuring its ability to displace [3H]-(+)-pentazocine from guinea pig brai...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M1 was determined by measuring its ability to displace [3H]-Pirenzepine from bovi...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M1 was determined by measuring its ability to displace [3H]-Pirenzepine from bovi...More data for this Ligand-Target Pair
Affinity DataKi: 36nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M1 was determined by measuring its ability to displace [3H]-Pirenzepine from bovi...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 39nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 39nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 41nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 59nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Homo sapiens (Human))
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 62.1nMAssay Description:In vitro binding affinity towards Sigma receptor type 1 was determined by measuring its ability to displace [3H]-(+)-pentazocine from guinea pig brai...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 80nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M3 was determined by measuring its ability to displace [3H]-N-methyl- scopolamine...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 85nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 160nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M3 was determined by measuring its ability to displace [3H]-N-methyl- scopolamine...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: 163nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M3 was determined by measuring its ability to displace [3H]-N-methyl- scopolamine...More data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:In vitro binding affinity towards Sigma receptor type 2 was determined by measuring its ability to displace [3H]-1,3-di-o-tolylguanidine from rat liv...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M3 was determined by measuring its ability to displace [3H]-N-methyl- scopolamine...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:In vitro binding affinity towards Muscarinic acetylcholine receptor M3 was determined by measuring its ability to displace [3H]-N-methyl- scopolamine...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor(Cavia porcellus)
National Institutes of Health
Curated by ChEMBL
National Institutes of Health
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:In vitro binding affinity against guinea pig ileum using [3H]-N-methylscopolamineMore data for this Ligand-Target Pair
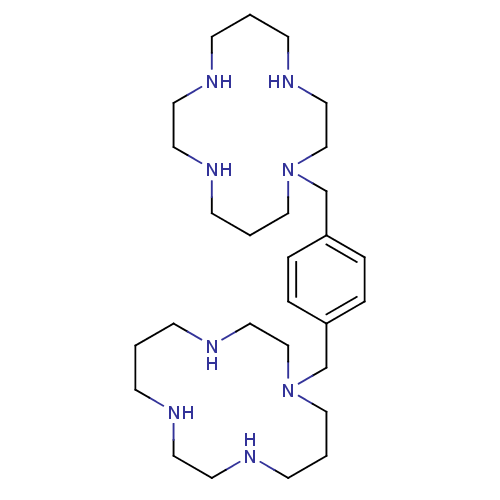
TargetC-X-C chemokine receptor type 4(Homo sapiens (Human))
National Institute of Biomedical Imaging and Bioengineering
Curated by ChEMBL
National Institute of Biomedical Imaging and Bioengineering
Curated by ChEMBL
Affinity DataIC50: 27.4nMAssay Description:Antagonist activity at CXCR4 in human Jurkat cells assessed as inhibition of SDF1-induced cell migrationMore data for this Ligand-Target Pair