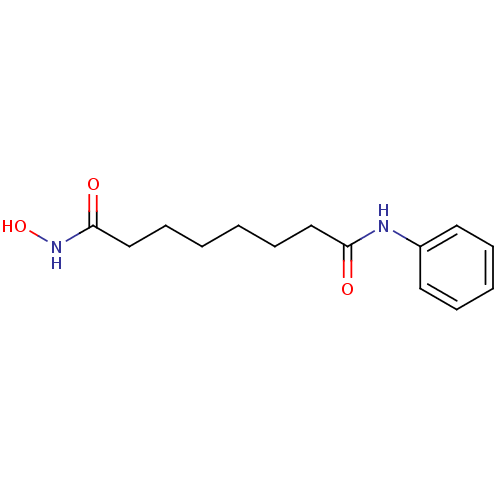
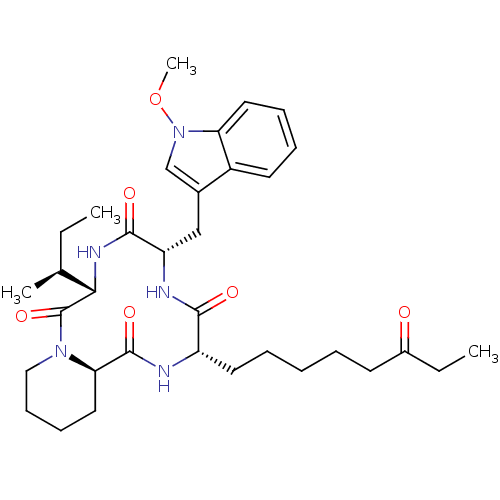
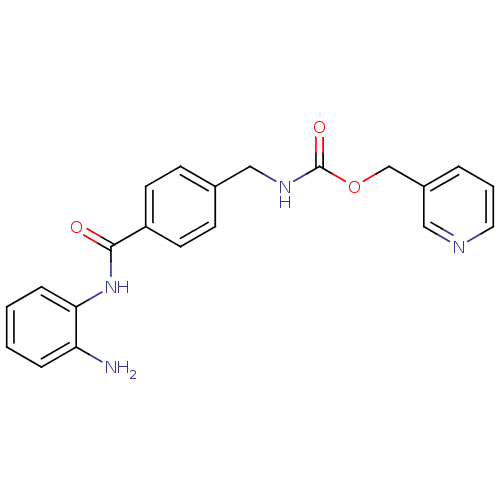
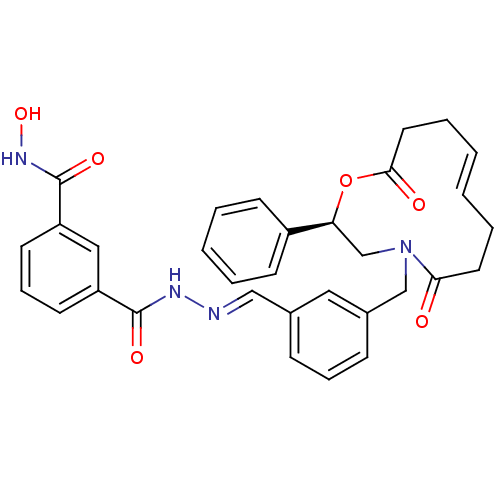
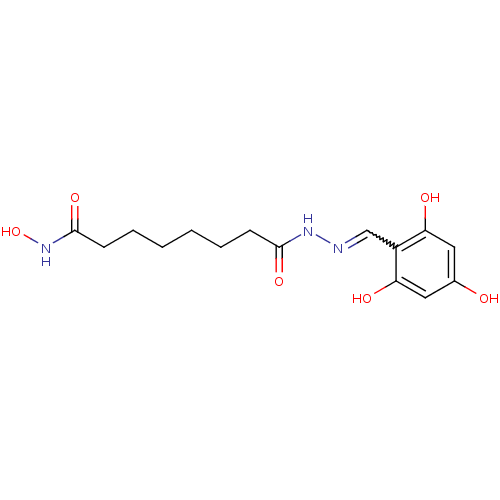
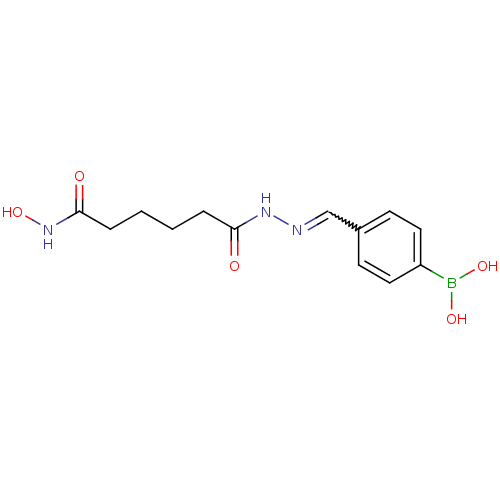
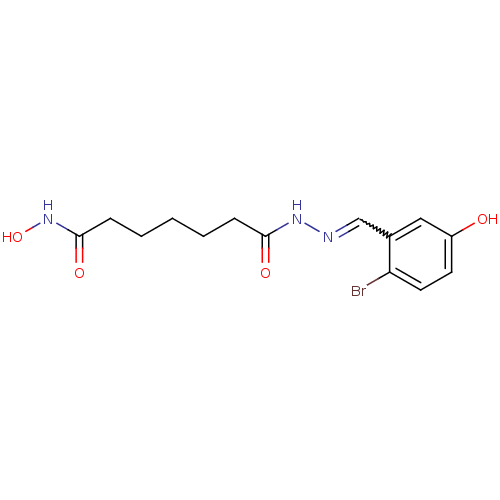
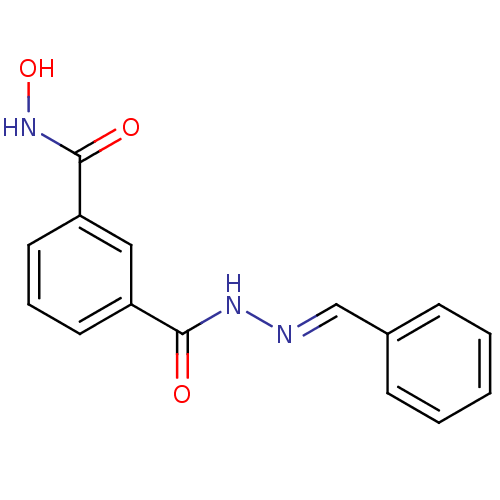
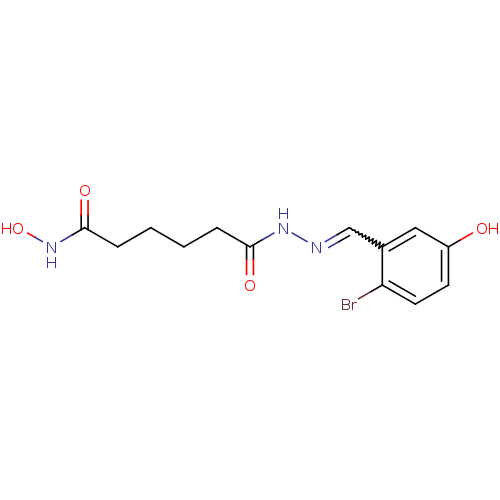
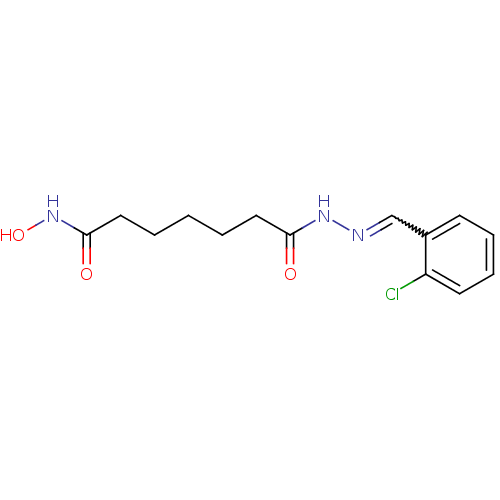
Affinity DataKi: 21nM ΔG°: -43.4kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
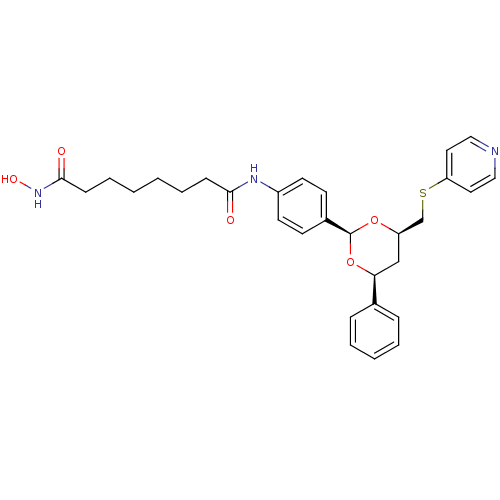
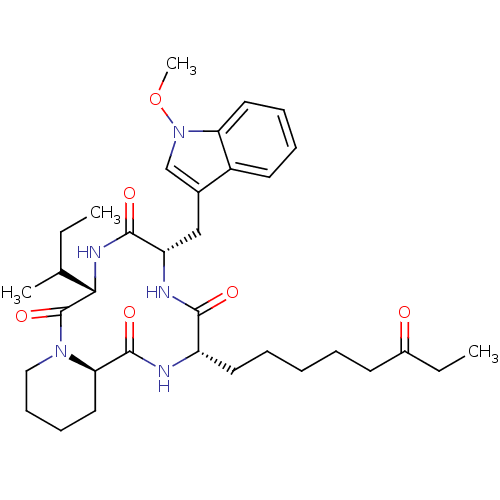
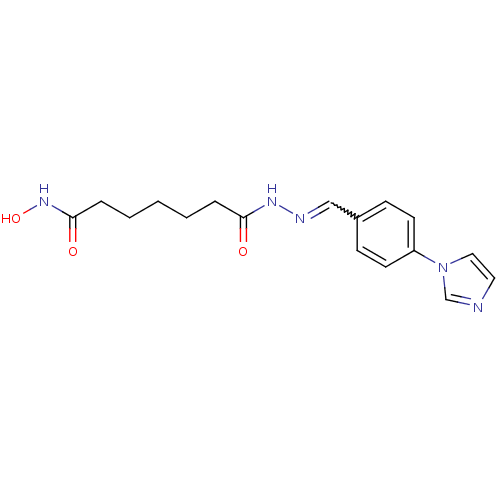
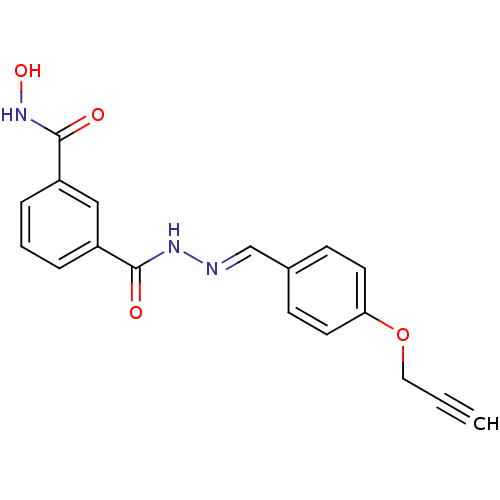
Affinity DataKi: 28nM ΔG°: -42.7kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
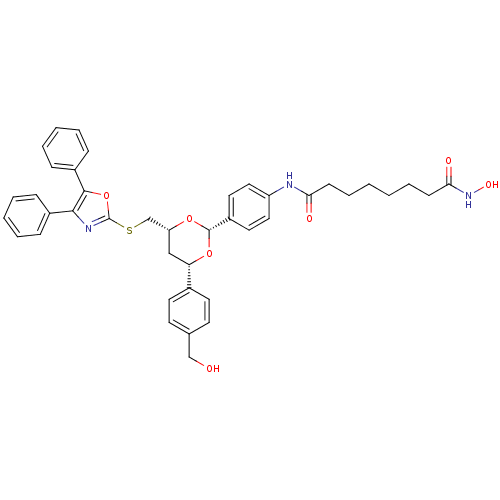
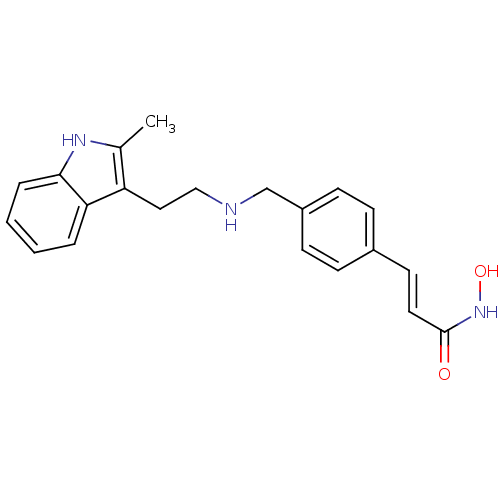
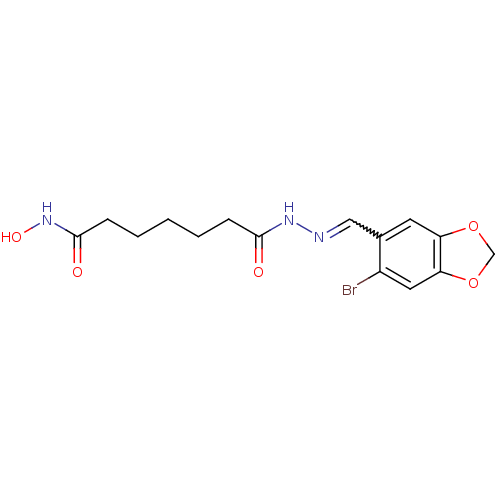
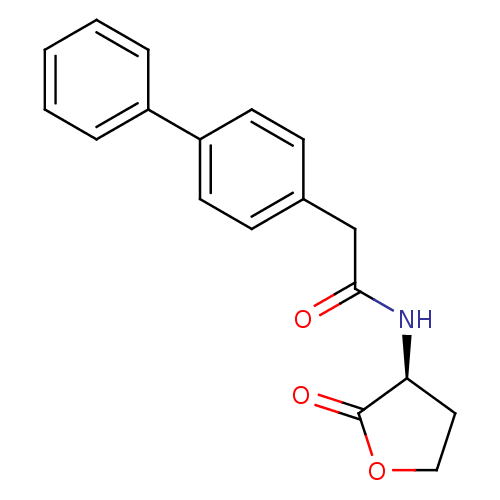
Affinity DataKi: 48nM ΔG°: -41.4kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
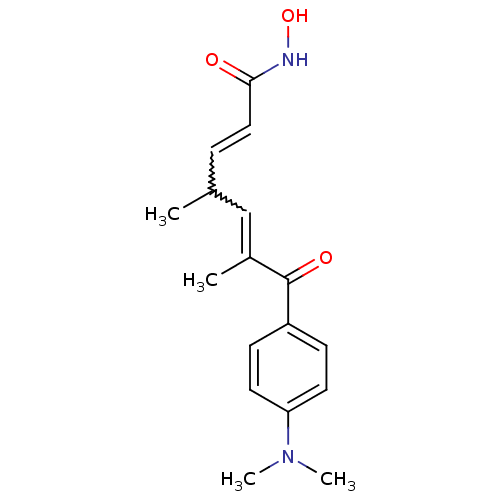
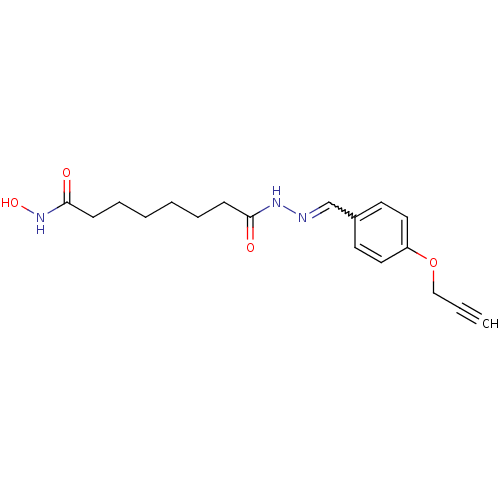
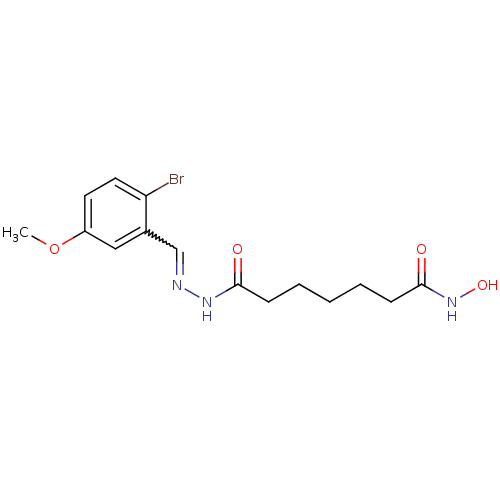
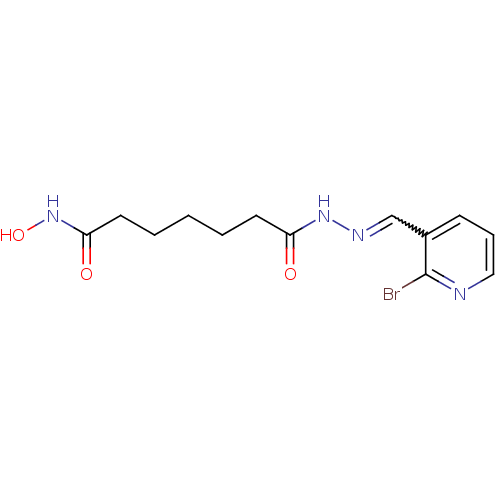
Affinity DataKi: 88nM ΔG°: -39.9kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 142nM ΔG°: -38.7kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 995nM ΔG°: -33.9kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM ΔG°: -32.2kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Target InfoPDBMMDBNCI pathwayReactome pathwayKEGG
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
UniProtKB/SwissProt
B.MOADDrugBankantibodypediaGoogleScholar
In DepthDetails
Article
BindingDB Entry DOI: 10.7270/Q2NC5ZG2PubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)
BindingDB Entry DOI: 10.7270/Q2NC5ZG2PubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)Affinity DataKi: 6.10E+3nM ΔG°: -29.5kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataKi: 6.30E+3nM ΔG°: -29.4kJ/molepH: 7.4 T: 2°CAssay Description:To assess the effect of test compounds on histone deacetylase enzyme function in Vitro, a fluorometric assay was performed using HDAC, which incubate...More data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMAssay Description:Inhibition of HDAC1More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibition of HDAC3More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibition of HDAC3More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition of HDAC1More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Inhibition of full length HDAC2 assessed as 7-amino-4-methylcoumarin release from fluorophore conjugated substrate after 5 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetHistone deacetylase 3/Nuclear receptor corepressor 2 (HDAC3/NCoR2)(Homo sapiens (Human))
Howard Hughes Medical Institute
Curated by ChEMBL
Howard Hughes Medical Institute
Curated by ChEMBL
Affinity DataIC50: 3.10nMAssay Description:Inhibition of full length HDAC3/NCoR2 assessed as 7-amino-4-methylcoumarin release from fluorophore conjugated substrate after 5 mins by fluorescence...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of full length HDAC8 assessed as 7-amino-4-methylcoumarin release from fluorophore conjugated substrate after 5 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of full length HDAC8 assessed as 7-amino-4-methylcoumarin release from fluorophore conjugated substrate after 5 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetQuorum-sensing transcriptional repressor QscR(Pseudomonas aeruginosa)
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Antagonist activity at Pseudomonas aeruginosa QscR expressed in Escherichia coli assessed as inhibition of dodecanoyl homoserine lactone induced prod...More data for this Ligand-Target Pair
TargetHistone deacetylase 3/Nuclear receptor corepressor 2 (HDAC3/NCoR2)(Homo sapiens (Human))
Howard Hughes Medical Institute
Curated by ChEMBL
Howard Hughes Medical Institute
Curated by ChEMBL
Affinity DataIC50: 34nMAssay Description:Inhibition of full length HDAC3/NCoR2 assessed as 7-amino-4-methylcoumarin release from fluorophore conjugated substrate after 5 mins by fluorescence...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 43nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 45nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMAssay Description:Inhibition of full length HDAC8 assessed as 7-amino-4-methylcoumarin release from fluorophore conjugated substrate after 5 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 66nMAssay Description:Inhibition of full length HDAC2 assessed as 7-amino-4-methylcoumarin release from fluorophore conjugated substrate after 5 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 89nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMpH: 7.5 T: 2°CAssay Description:Recombinant pfHDAC-1 was assayed with substrate in the presence of test compound. The substrate concentration was kept constant at 125 uM while the c...More data for this Ligand-Target Pair