Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Cyclin-dependent kinase 2/G1/S-specific cyclin-E1
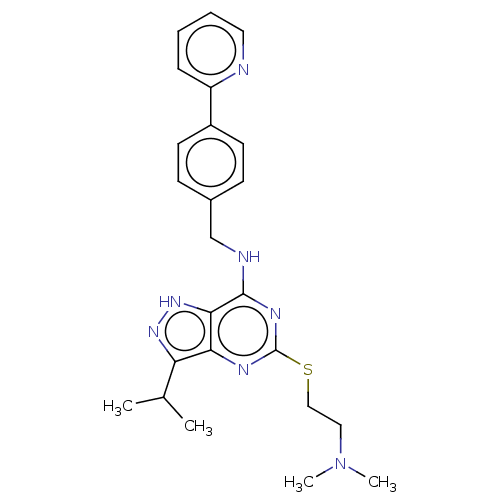
Ligand
BDBM50521564
Substrate
n/a
Meas. Tech.
ChEMBL_1884560 (CHEMBL4386142)
IC50
77±n/a nM
Citation
 Jorda, R; Havl�?ek, L; ?turc, A; Tu?kov�, D; Daumov�, L; Alam, M; ?kerlov�, J; Nekardov�, M; Pe?ina, M; Posp�?il, T; ?irok�, J; Urb�nek, L; Pachl, P; ?ez�?ov�, P; Strnad, M; Klener, P; Kry?tof, V 3,5,7-Substituted Pyrazolo[4,3- d]pyrimidine Inhibitors of Cyclin-Dependent Kinases and Their Evaluation in Lymphoma Models. J Med Chem 62:4606-4623 (2019) [PubMed] Article
Jorda, R; Havl�?ek, L; ?turc, A; Tu?kov�, D; Daumov�, L; Alam, M; ?kerlov�, J; Nekardov�, M; Pe?ina, M; Posp�?il, T; ?irok�, J; Urb�nek, L; Pachl, P; ?ez�?ov�, P; Strnad, M; Klener, P; Kry?tof, V 3,5,7-Substituted Pyrazolo[4,3- d]pyrimidine Inhibitors of Cyclin-Dependent Kinases and Their Evaluation in Lymphoma Models. J Med Chem 62:4606-4623 (2019) [PubMed] ArticleMore Info.:
Target
Name:
Cyclin-dependent kinase 2/G1/S-specific cyclin-E1
Synonyms:
CDK2/CycE | CDK2/Cyclin E | CDK2/E | CDK2/E1 | Cyclin-Dependent Kinase 2 (CDK2) | Cyclin-dependent kinase 2/G1/S-specific cyclin E1 | Cyclin-dependent kinase 2/cyclin E1
Type:
Protein
Mol. Mass.:
n/a
Description:
n/a
Components:
This complex has 2 components.
Component 1
Name:
Cyclin-dependent kinase 2
Synonyms:
CDK2 | CDK2-Kinase | CDK2_HUMAN | CDKN2 | Cell division protein kinase 2 | Cyclin-dependent kinase 2 (CDK2) | Protein cereblon/Cyclin-dependent kinase 2 | p33 protein kinase
Type:
Enzyme Subunit
Mol. Mass.:
33938.17
Organism:
Homo sapiens (Human)
Description:
P24941
Residue:
298
Sequence:
MENFQKVEKIGEGTYGVVYKARNKLTGEVVALKKIRLDTETEGVPSTAIREISLLKELNHPNIVKLLDVIHTENKLYLVFEFLHQDLKKFMDASALTGIPLPLIKSYLFQLLQGLAFCHSHRVLHRDLKPQNLLINTEGAIKLADFGLARAFGVPVRTYTHEVVTLWYRAPEILLGCKYYSTAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVWPGVTSMPDYKPSFPKWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFFQDVTKPVPHLRL
Component 2
Name:
G1/S-specific cyclin-E1
Synonyms:
CCNE | CCNE1 | CCNE1_HUMAN
Type:
Enzyme Subunit
Mol. Mass.:
47073.17
Organism:
Homo sapiens (Human)
Description:
n/a
Residue:
410
Sequence:
MPRERRERDAKERDTMKEDGGAEFSARSRKRKANVTVFLQDPDEEMAKIDRTARDQCGSQPWDNNAVCADPCSLIPTPDKEDDDRVYPNSTCKPRIIAPSRGSPLPVLSWANREEVWKIMLNKEKTYLRDQHFLEQHPLLQPKMRAILLDWLMEVCEVYKLHRETFYLAQDFFDRYMATQENVVKTLLQLIGISSLFIAAKLEEIYPPKLHQFAYVTDGACSGDEILTMELMIMKALKWRLSPLTIVSWLNVYMQVAYLNDLHEVLLPQYPQQIFIQIAELLDLCVLDVDCLEFPYGILAASALYHFSSSELMQKVSGYQWCDIENCVKWMVPFAMVIRETGSSKLKHFRGVADEDAHNIQTHRDSLDLLDKARAKKAMLSEQNRASPLPSGLLTPPQSGKKQSSGPEMA