Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Transcription intermediary factor 1-alpha
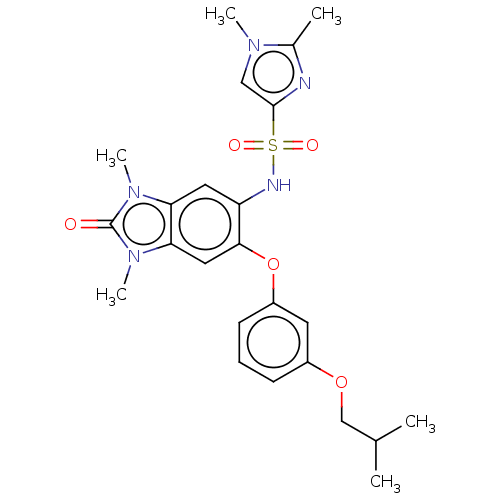
Ligand
BDBM50150808
Substrate
n/a
Meas. Tech.
ChEMBL_1560318 (CHEMBL3779310)
IC50
57±n/a nM
Citation
 Palmer, WS; Poncet-Montange, G; Liu, G; Petrocchi, A; Reyna, N; Subramanian, G; Theroff, J; Yau, A; Kost-Alimova, M; Bardenhagen, JP; Leo, E; Shepard, HE; Tieu, TN; Shi, X; Zhan, Y; Zhao, S; Barton, MC; Draetta, G; Toniatti, C; Jones, P; Geck Do, M; Andersen, JN Structure-Guided Design of IACS-9571, a Selective High-Affinity Dual TRIM24-BRPF1 Bromodomain Inhibitor. J Med Chem 59:1440-54 (2016) [PubMed] Article
Palmer, WS; Poncet-Montange, G; Liu, G; Petrocchi, A; Reyna, N; Subramanian, G; Theroff, J; Yau, A; Kost-Alimova, M; Bardenhagen, JP; Leo, E; Shepard, HE; Tieu, TN; Shi, X; Zhan, Y; Zhao, S; Barton, MC; Draetta, G; Toniatti, C; Jones, P; Geck Do, M; Andersen, JN Structure-Guided Design of IACS-9571, a Selective High-Affinity Dual TRIM24-BRPF1 Bromodomain Inhibitor. J Med Chem 59:1440-54 (2016) [PubMed] ArticleMore Info.:
Target
Name:
Transcription intermediary factor 1-alpha
Synonyms:
6.3.2.- | E3 ubiquitin-protein ligase TRIM24 | RING finger protein 82 | RNF82 | RNF82 | TIF1 | TIF1-alpha | TIF1A | TIF1A_HUMAN | TRIM24 | Transcription intermediary factor 1-alpha | Tripartite motif-containing protein 24
Type:
PROTEIN
Mol. Mass.:
116839.55
Organism:
Homo sapiens (Human)
Description:
ChEMBL_107998
Residue:
1050
Sequence:
MEVAVEKAVAAAAAASAAASGGPSAAPSGENEAESRQGPDSERGGEAARLNLLDTCAVCHQNIQSRAPKLLPCLHSFCQRCLPAPQRYLMLPAPMLGSAETPPPVPAPGSPVSGSSPFATQVGVIRCPVCSQECAERHIIDNFFVKDTTEVPSSTVEKSNQVCTSCEDNAEANGFCVECVEWLCKTCIRAHQRVKFTKDHTVRQKEEVSPEAVGVTSQRPVFCPFHKKEQLKLYCETCDKLTCRDCQLLEHKEHRYQFIEEAFQNQKVIIDTLITKLMEKTKYIKFTGNQIQNRIIEVNQNQKQVEQDIKVAIFTLMVEINKKGKALLHQLESLAKDHRMKLMQQQQEVAGLSKQLEHVMHFSKWAVSSGSSTALLYSKRLITYRLRHLLRARCDASPVTNNTIQFHCDPSFWAQNIINLGSLVIEDKESQPQMPKQNPVVEQNSQPPSGLSSNQLSKFPTQISLAQLRLQHMQQQVMAQRQQVQRRPAPVGLPNPRMQGPIQQPSISHQQPPPRLINFQNHSPKPNGPVLPPHPQQLRYPPNQNIPRQAIKPNPLQMAFLAQQAIKQWQISSGQGTPSTTNSTSSTPSSPTITSAAGYDGKAFGSPMIDLSSPVGGSYNLPSLPDIDCSSTIMLDNIVRKDTNIDHGQPRPPSNRTVQSPNSSVPSPGLAGPVTMTSVHPPIRSPSASSVGSRGSSGSSSKPAGADSTHKVPVVMLEPIRIKQENSGPPENYDFPVVIVKQESDEESRPQNANYPRSILTSLLLNSSQSSTSEETVLRSDAPDSTGDQPGLHQDNSSNGKSEWLDPSQKSPLHVGETRKEDDPNEDWCAVCQNGGELLCCEKCPKVFHLSCHVPTLTNFPSGEWICTFCRDLSKPEVEYDCDAPSHNSEKKKTEGLVKLTPIDKRKCERLLLFLYCHEMSLAFQDPVPLTVPDYYKIIKNPMDLSTIKKRLQEDYSMYSKPEDFVADFRLIFQNCAEFNEPDSEVANAGIKLENYFEELLKNLYPEKRFPKPEFRNESEDNKFSDDSDDDFVQPRKKRLKSIEERQLLK