Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Macrophage colony-stimulating factor 1 receptor
Ligand
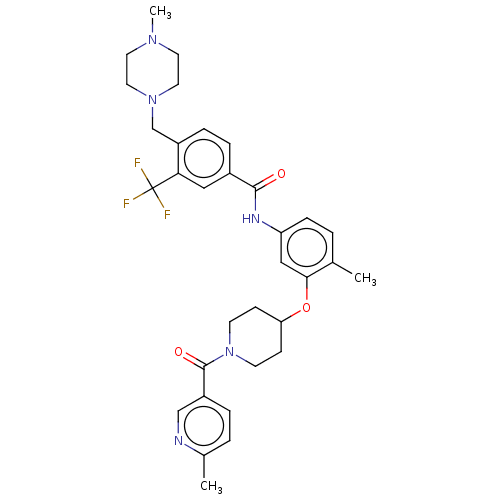
BDBM50211223
Substrate
n/a
Meas. Tech.
ChEMBL_1636087 (CHEMBL3878985)
IC50
227±n/a nM
Citation
 Wang, Q; Liu, F; Wang, B; Zou, F; Qi, Z; Chen, C; Yu, K; Hu, C; Qi, S; Wang, W; Hu, Z; Liu, J; Wang, W; Wang, L; Liang, Q; Zhang, S; Ren, T; Liu, Q; Liu, J Discovery of 4-Methyl-N-(4-((4-methylpiperazin-1-yl)methyl)-3-(trifluoromethyl)phenyl)-3-((1-nicotinoylpiperidin-4-yl)oxy)benzamide (CHMFL-ABL/KIT-155) as a Novel Highly Potent Type II ABL/KIT Dual Kinase Inhibitor with a Distinct Hinge Binding. J Med Chem 60:273-289 (2017) [PubMed] Article
Wang, Q; Liu, F; Wang, B; Zou, F; Qi, Z; Chen, C; Yu, K; Hu, C; Qi, S; Wang, W; Hu, Z; Liu, J; Wang, W; Wang, L; Liang, Q; Zhang, S; Ren, T; Liu, Q; Liu, J Discovery of 4-Methyl-N-(4-((4-methylpiperazin-1-yl)methyl)-3-(trifluoromethyl)phenyl)-3-((1-nicotinoylpiperidin-4-yl)oxy)benzamide (CHMFL-ABL/KIT-155) as a Novel Highly Potent Type II ABL/KIT Dual Kinase Inhibitor with a Distinct Hinge Binding. J Med Chem 60:273-289 (2017) [PubMed] ArticleMore Info.:
Target
Name:
Macrophage colony-stimulating factor 1 receptor
Synonyms:
CSF1R | CSF1R_HUMAN | FMS | Fms proto-oncogene | Macrophage colony stimulating factor receptor | Macrophage colony-stimulating factor 1 receptor (FMS) | Macrophage colony-stimulating factor 1 receptor (c-FMS) | Macrophage colony-stimulating factor 1 receptor (cFMS) | Proto-oncogene c-Fms | Tyrosine Kinase CSF1-R | c-fms | cFMS
Type:
Protein
Mol. Mass.:
107979.48
Organism:
Homo sapiens (Human)
Description:
P07333
Residue:
972
Sequence:
MGPGVLLLLLVATAWHGQGIPVIEPSVPELVVKPGATVTLRCVGNGSVEWDGPPSPHWTLYSDGSSSILSTNNATFQNTGTYRCTEPGDPLGGSAAIHLYVKDPARPWNVLAQEVVVFEDQDALLPCLLTDPVLEAGVSLVRVRGRPLMRHTNYSFSPWHGFTIHRAKFIQSQDYQCSALMGGRKVMSISIRLKVQKVIPGPPALTLVPAELVRIRGEAAQIVCSASSVDVNFDVFLQHNNTKLAIPQQSDFHNNRYQKVLTLNLDQVDFQHAGNYSCVASNVQGKHSTSMFFRVVESAYLNLSSEQNLIQEVTVGEGLNLKVMVEAYPGLQGFNWTYLGPFSDHQPEPKLANATTKDTYRHTFTLSLPRLKPSEAGRYSFLARNPGGWRALTFELTLRYPPEVSVIWTFINGSGTLLCAASGYPQPNVTWLQCSGHTDRCDEAQVLQVWDDPYPEVLSQEPFHKVTVQSLLTVETLEHNQTYECRAHNSVGSGSWAFIPISAGAHTHPPDEFLFTPVVVACMSIMALLLLLLLLLLYKYKQKPKYQVRWKIIESYEGNSYTFIDPTQLPYNEKWEFPRNNLQFGKTLGAGAFGKVVEATAFGLGKEDAVLKVAVKMLKSTAHADEKEALMSELKIMSHLGQHENIVNLLGACTHGGPVLVITEYCCYGDLLNFLRRKAEAMLGPSLSPGQDPEGGVDYKNIHLEKKYVRRDSGFSSQGVDTYVEMRPVSTSSNDSFSEQDLDKEDGRPLELRDLLHFSSQVAQGMAFLASKNCIHRDVAARNVLLTNGHVAKIGDFGLARDIMNDSNYIVKGNARLPVKWMAPESIFDCVYTVQSDVWSYGILLWEIFSLGLNPYPGILVNSKFYKLVKDGYQMAQPAFAPKNIYSIMQACWALEPTHRPTFQQICSFLQEQAQEDRRERDYTNLPSSSRSGGSGSSSSELEEESSSEHLTCCEQGDIAQPLLQPNNYQFC