Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Histone-lysine N-methyltransferase EHMT2
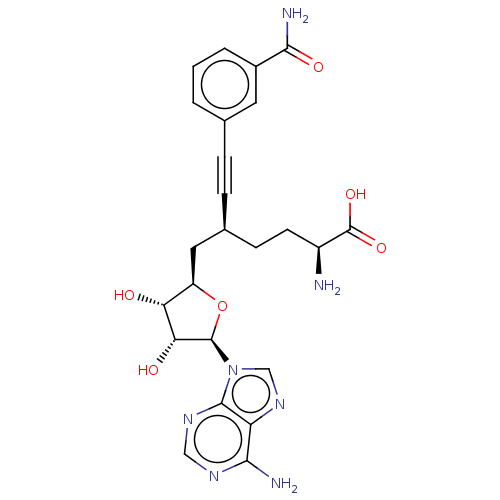
Ligand
BDBM50530712
Substrate
n/a
Meas. Tech.
ChEMBL_1913771 (CHEMBL4416354)
IC50
>10000±n/a nM
Citation
 Policarpo, RL; Decultot, L; May, E; Kuzmi?, P; Carlson, S; Huang, D; Chu, V; Wright, BA; Dhakshinamoorthy, S; Kannt, A; Rani, S; Dittakavi, S; Panarese, JD; Gaudet, R; Shair, MD High-Affinity Alkynyl Bisubstrate Inhibitors of Nicotinamide J Med Chem 62:9837-9873 (2019) [PubMed] Article
Policarpo, RL; Decultot, L; May, E; Kuzmi?, P; Carlson, S; Huang, D; Chu, V; Wright, BA; Dhakshinamoorthy, S; Kannt, A; Rani, S; Dittakavi, S; Panarese, JD; Gaudet, R; Shair, MD High-Affinity Alkynyl Bisubstrate Inhibitors of Nicotinamide J Med Chem 62:9837-9873 (2019) [PubMed] ArticleMore Info.:
Target
Name:
Histone-lysine N-methyltransferase EHMT2
Synonyms:
BAT8 | C6orf30 | EHMT2 | EHMT2_HUMAN | G9A | G9a histone methyltransferase (G9A) | Histone-lysine N-methyltransferase EHMT1/EHMT2 | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 | KMT1C | NG36 | Protein G9a (G9a)
Type:
Enzyme
Mol. Mass.:
132339.87
Organism:
Homo sapiens (Human)
Description:
Q96KQ7
Residue:
1210
Sequence:
MAAAAGAAAAAAAEGEAPAEMGALLLEKETRGATERVHGSLGDTPRSEETLPKATPDSLEPAGPSSPASVTVTVGDEGADTPVGATPLIGDESENLEGDGDLRGGRILLGHATKSFPSSPSKGGSCPSRAKMSMTGAGKSPPSVQSLAMRLLSMPGAQGAAAAGSEPPPATTSPEGQPKVHRARKTMSKPGNGQPPVPEKRPPEIQHFRMSDDVHSLGKVTSDLAKRRKLNSGGGLSEELGSARRSGEVTLTKGDPGSLEEWETVVGDDFSLYYDSYSVDERVDSDSKSEVEALTEQLSEEEEEEEEEEEEEEEEEEEEEEEEDEESGNQSDRSGSSGRRKAKKKWRKDSPWVKPSRKRRKREPPRAKEPRGVNGVGSSGPSEYMEVPLGSLELPSEGTLSPNHAGVSNDTSSLETERGFEELPLCSCRMEAPKIDRISERAGHKCMATESVDGELSGCNAAILKRETMRPSSRVALMVLCETHRARMVKHHCCPGCGYFCTAGTFLECHPDFRVAHRFHKACVSQLNGMVFCPHCGEDASEAQEVTIPRGDGVTPPAGTAAPAPPPLSQDVPGRADTSQPSARMRGHGEPRRPPCDPLADTIDSSGPSLTLPNGGCLSAVGLPLGPGREALEKALVIQESERRKKLRFHPRQLYLSVKQGELQKVILMLLDNLDPNFQSDQQSKRTPLHAAAQKGSVEICHVLLQAGANINAVDKQQRTPLMEAVVNNHLEVARYMVQRGGCVYSKEEDGSTCLHHAAKIGNLEMVSLLLSTGQVDVNAQDSGGWTPIIWAAEHKHIEVIRMLLTRGADVTLTDNEENICLHWASFTGSAAIAEVLLNARCDLHAVNYHGDTPLHIAARESYHDCVLLFLSRGANPELRNKEGDTAWDLTPERSDVWFALQLNRKLRLGVGNRAIRTEKIICRDVARGYENVPIPCVNGVDGEPCPEDYKYISENCETSTMNIDRNITHLQHCTCVDDCSSSNCLCGQLSIRCWYDKDGRLLQEFNKIEPPLIFECNQACSCWRNCKNRVVQSGIKVRLQLYRTAKMGWGVRALQTIPQGTFICEYVGELISDAEADVREDDSYLFDLDNKDGEVYCIDARYYGNISRFINHLCDPNIIPVRVFMLHQDLRFPRIAFFSSRDIRTGEELGFDYGDRFWDIKSKYFTCQCGSEKCKHSAEAIALEQSRLARLDPHPELLPELGSLPPVNT