Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Histone-lysine N-methyltransferase EHMT2
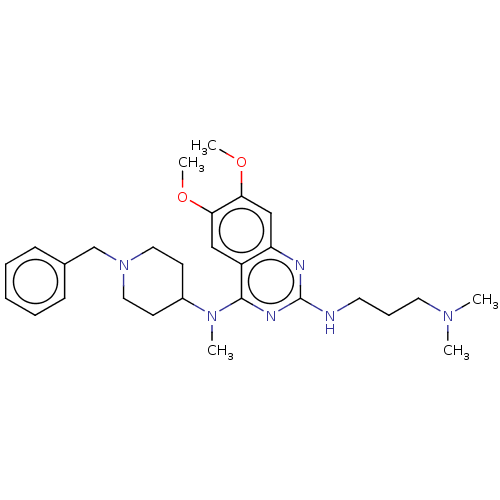
Ligand
BDBM50594087
Substrate
n/a
Meas. Tech.
ChEMBL_2210856 (CHEMBL5123805)
Ki
>100000±n/a nM
Citation
 Menna, M; Fiorentino, F; Marrocco, B; Lucidi, A; Tomassi, S; Cilli, D; Romanenghi, M; Cassandri, M; Pomella, S; Pezzella, M; Del Bufalo, D; Zeya Ansari, MS; Toma?evi?, N; Mladenovi?, M; Viviano, M; Sbardella, G; Rota, R; Trisciuoglio, D; Minucci, S; Mattevi, A; Rotili, D; Mai, A Novel non-covalent LSD1 inhibitors endowed with anticancer effects in leukemia and solid tumor cellular models. Eur J Med Chem 237:0 (2022) [PubMed] Article
Menna, M; Fiorentino, F; Marrocco, B; Lucidi, A; Tomassi, S; Cilli, D; Romanenghi, M; Cassandri, M; Pomella, S; Pezzella, M; Del Bufalo, D; Zeya Ansari, MS; Toma?evi?, N; Mladenovi?, M; Viviano, M; Sbardella, G; Rota, R; Trisciuoglio, D; Minucci, S; Mattevi, A; Rotili, D; Mai, A Novel non-covalent LSD1 inhibitors endowed with anticancer effects in leukemia and solid tumor cellular models. Eur J Med Chem 237:0 (2022) [PubMed] ArticleMore Info.:
Target
Name:
Histone-lysine N-methyltransferase EHMT2
Synonyms:
BAT8 | C6orf30 | EHMT2 | EHMT2_HUMAN | G9A | G9a histone methyltransferase (G9A) | Histone-lysine N-methyltransferase EHMT1/EHMT2 | Histone-lysine N-methyltransferase, H3 lysine-9 specific 3 | KMT1C | NG36 | Protein G9a (G9a)
Type:
Enzyme
Mol. Mass.:
132339.87
Organism:
Homo sapiens (Human)
Description:
Q96KQ7
Residue:
1210
Sequence:
MAAAAGAAAAAAAEGEAPAEMGALLLEKETRGATERVHGSLGDTPRSEETLPKATPDSLEPAGPSSPASVTVTVGDEGADTPVGATPLIGDESENLEGDGDLRGGRILLGHATKSFPSSPSKGGSCPSRAKMSMTGAGKSPPSVQSLAMRLLSMPGAQGAAAAGSEPPPATTSPEGQPKVHRARKTMSKPGNGQPPVPEKRPPEIQHFRMSDDVHSLGKVTSDLAKRRKLNSGGGLSEELGSARRSGEVTLTKGDPGSLEEWETVVGDDFSLYYDSYSVDERVDSDSKSEVEALTEQLSEEEEEEEEEEEEEEEEEEEEEEEEDEESGNQSDRSGSSGRRKAKKKWRKDSPWVKPSRKRRKREPPRAKEPRGVNGVGSSGPSEYMEVPLGSLELPSEGTLSPNHAGVSNDTSSLETERGFEELPLCSCRMEAPKIDRISERAGHKCMATESVDGELSGCNAAILKRETMRPSSRVALMVLCETHRARMVKHHCCPGCGYFCTAGTFLECHPDFRVAHRFHKACVSQLNGMVFCPHCGEDASEAQEVTIPRGDGVTPPAGTAAPAPPPLSQDVPGRADTSQPSARMRGHGEPRRPPCDPLADTIDSSGPSLTLPNGGCLSAVGLPLGPGREALEKALVIQESERRKKLRFHPRQLYLSVKQGELQKVILMLLDNLDPNFQSDQQSKRTPLHAAAQKGSVEICHVLLQAGANINAVDKQQRTPLMEAVVNNHLEVARYMVQRGGCVYSKEEDGSTCLHHAAKIGNLEMVSLLLSTGQVDVNAQDSGGWTPIIWAAEHKHIEVIRMLLTRGADVTLTDNEENICLHWASFTGSAAIAEVLLNARCDLHAVNYHGDTPLHIAARESYHDCVLLFLSRGANPELRNKEGDTAWDLTPERSDVWFALQLNRKLRLGVGNRAIRTEKIICRDVARGYENVPIPCVNGVDGEPCPEDYKYISENCETSTMNIDRNITHLQHCTCVDDCSSSNCLCGQLSIRCWYDKDGRLLQEFNKIEPPLIFECNQACSCWRNCKNRVVQSGIKVRLQLYRTAKMGWGVRALQTIPQGTFICEYVGELISDAEADVREDDSYLFDLDNKDGEVYCIDARYYGNISRFINHLCDPNIIPVRVFMLHQDLRFPRIAFFSSRDIRTGEELGFDYGDRFWDIKSKYFTCQCGSEKCKHSAEAIALEQSRLARLDPHPELLPELGSLPPVNT