Reaction Details  Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
S-adenosylmethionine synthase isoform type-1/type-2
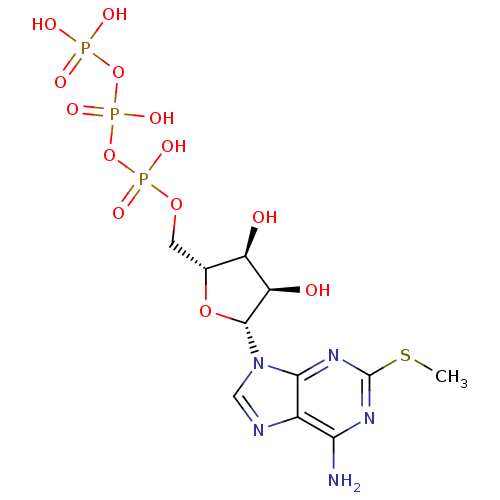
Ligand
BDBM50118232
Substrate
n/a
Meas. Tech.
ChEMBL_104961 (CHEMBL878217)
Ki
110000±n/a nM
Citation
 Kappler, F; Hai, TT; Hampton, A Isozyme-specific enzyme inhibitors. 10. Adenosine 5'-triphosphate derivatives as substrates or inhibitors of methionine adenosyltransferases of rat normal and hepatoma tissues. J Med Chem 29:318-22 (1986) [PubMed]
Kappler, F; Hai, TT; Hampton, A Isozyme-specific enzyme inhibitors. 10. Adenosine 5'-triphosphate derivatives as substrates or inhibitors of methionine adenosyltransferases of rat normal and hepatoma tissues. J Med Chem 29:318-22 (1986) [PubMed]More Info.:
Target
Name:
S-adenosylmethionine synthase isoform type-1/type-2
Synonyms:
S-adenosylmethionine synthetase (MAT 1 and MAT 2)
Type:
n/a
Mol. Mass.:
n/a
Description:
ASSAY_ID of ChEMBL is 104961
Components:
This complex has 2 components.
Component 1
Name:
S-adenosylmethionine synthase isoform type-1
Synonyms:
AdoMet synthetase 1 | Ams1 | MAT 1 | MAT-I/III | METK1_RAT | Mat1a | Methionine adenosyltransferase 1 | Methionine adenosyltransferase I/III | S-adenosylmethionine synthetase (MAT 1 and MAT 2) | S-adenosylmethionine synthetase alpha and beta forms | S-adenosylmethionine synthetase isoform type-1
Type:
PROTEIN
Mol. Mass.:
43692.87
Organism:
Rattus norvegicus
Description:
ChEMBL_11027
Residue:
397
Sequence:
MNGPVDGLCDHSLSEEGAFMFTSESVGEGHPDKICDQISDAVLDAHLKQDPNAKVACETVCKTGMVLLCGEITSMAMIDYQRVVRDTIKHIGYDDSAKGFDFKTCNVLVALEQQSPDIAQCVHLDRNEEDVGAGDQGLMFGYATDETEECMPLTIVLAHKLNTRMADLRRSGVLPWLRPDSKTQVTVQYVQDNGAVIPVRVHTIVISVQHNEDITLEAMREALKEQVIKAVVPAKYLDEDTIYHLQPSGRFVIGGPQGDAGVTGRKIIVDTYGGWGAHGGGAFSGKDYTKVDRSAAYAARWVAKSLVKAGLCRRVLVQVSYAIGVAEPLSISIFTYGTSKKTERDELLEVVNKNFDLRPGVIVRDLDLKKPIYQKTACYGHFGRSEFPWEVPKKLVF
Component 2
Name:
S-adenosylmethionine synthase isoform type-2
Synonyms:
AdoMet synthetase 2 | Ams2 | MAT 2 | MAT-II | METK2_RAT | Mat2a | Methionine adenosyltransferase 2 | Methionine adenosyltransferase II | S-Adenosylmethionine Synthase (AdoMet) | S-adenosylmethionine synthetase (MAT 1 and MAT 2) | S-adenosylmethionine synthetase gamma form | S-adenosylmethionine synthetase isoform type-2
Type:
PROTEIN
Mol. Mass.:
43713.68
Organism:
Rattus norvegicus
Description:
ChEMBL_105118
Residue:
395
Sequence:
MNGQLNGFHEAFIEEGTFLFTSESVGEGHPDKICDQINDAVLDAHLQQDPDAKVACETVAKTGMILLAGEITSRAAIDYQKVVREAIKHIGYDDSSKGFDYKTCNVLVALEQQSPDIAQGVHLDRNEEDIGAGDQGLMFGYATDETEECMPLTIVLAHKLNAKLAELRRNGTLPWLRPDSKTQVTVQYMQDRGAVIPIRVHTIVISVQHDEEVCLDEMRDALKEKLIKAVVPAKYLDEDTIYHLQPSGRFVIGGPQGDAGLTGRKIIVDTYGGWGAHGGGAFSGKDYTKVDRSAAYAARWVAKSLVKGGLCRRVLVQVSYAIGVSHPLSISIFHYGTSQKSERELLEIVKNNFDLRPGVIVRDLDLKKPIYQRTAAYGHFGRDSFPWEVPKKLKY