

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
BDBM10851 (4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihydroxy-hexahydro-1,3,5,2-furo[3,2-d][1,3,2]dioxaphosphinin-2-one::(4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)tetrahydro-4H-furo[3,2-d][1,3,2]dioxaphosphinine-2,7-diol 2-oxide::ADENOSINE-3 ,5 -CYCLIC-MONOPHOSPHATE::Cyclic adenosine monophosphate::cAMP::cid_23902375
SMILES: Nc1ncnc2n(cnc12)[C@@H]1O[C@@H]2COP(O)(=O)O[C@H]2[C@H]1O
InChI Key: InChIKey=IVOMOUWHDPKRLL-KQYNXXCUSA-N
PDB links: 33 PDB IDs match this monomer. 65 PDB IDs contain inhibitors having a similarity of 90% to this monomer.
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kcal/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| cyclic AMP receptor protein (Vibrio cholerae O1 biovar El Tor str. N16961) | BDBM10851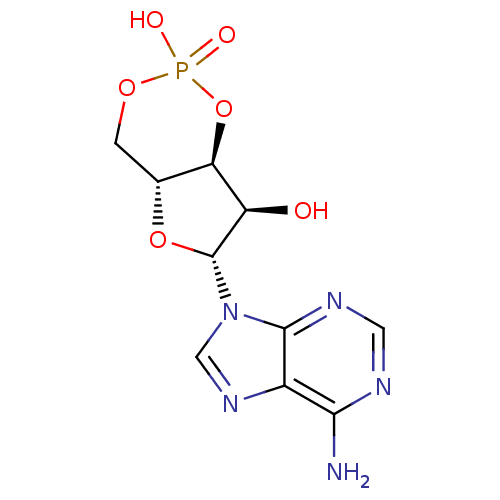 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | PDB MMDB KEGG UniProtKB/TrEMBL B.MOAD GoogleScholar AffyNet | Purchase KEGG PC cid PC sid PDB UniChem Similars | PDB PCBioAssay | n/a | n/a | 1.30E+5 | n/a | n/a | n/a | n/a | n/a | n/a |
Southern Research Specialized Biocontainment Screening Center Curated by PubChem BioAssay | PubChem Bioassay (2013) BindingDB Entry DOI: 10.7270/Q2HD7T8K | ||||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| Cell (A) | Syringe (B) | Cell Links | Syringe Links | Cell + Syr Links | ΔG° kcal/mole | -TΔS° kcal/mole | ΔH° kcal/mole | log K | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|
| Heat Shock Protein 90 (Hsp90) (Saccharomyces cerevisiae (Baker s yeast)) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -5.61 | 5.80 | -11.4 | 4.34 | 8 | 10 | |
University College London | J Mol Biol 392: 923-36 (2009) | |||||||||
| Heat Shock Protein 90 (Hsp90) (Saccharomyces cerevisiae (Baker s yeast)) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -5.40 | 6.07 | -11.5 | 4.08 | 8 | 15 | |
University College London | J Mol Biol 392: 923-36 (2009) | |||||||||
| Heat Shock Protein 90 (Hsp90) (Saccharomyces cerevisiae (Baker s yeast)) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -5.09 | 4.97 | -10.1 | 3.70 | 8 | 25 | |
University College London | J Mol Biol 392: 923-36 (2009) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -6.59 | -14.8 | 8.19 | 4.91 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -7.00 | -3.92 | -2.70 | 5.26 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -6.90 | -4.91 | -2.01 | 5.08 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -6.09 | -6.98 | 0.693 | 4.56 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -7.28 | -5.48 | -1.89 | 5.43 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -7.69 | -13.7 | 6.09 | 5.61 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -6.81 | -3.92 | -2.89 | 4.91 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -6.59 | -16.4 | 9.79 | 4.91 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -6.81 | -2.28 | -4.59 | 4.91 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -7.19 | -19.9 | 12.3 | 5.26 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -7.19 | -4.98 | -2.20 | 5.26 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||
| cAMP-activated global transcriptional regulator CRP (Escherichia coli) | BDBM10851 ((4aR,6R,7R,7aS)-6-(6-amino-9H-purin-9-yl)-2,7-dihy...) | GoogleScholar PDB | CHEBI KEGG PC cid PC sid PDB | -6.90 | -14.1 | 7.19 | 5.08 | 7.80 | 25 | |
Durham University | PLoS Biol 11: (2013) | |||||||||