| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Beta-secretase 1 |
|---|
| Ligand | BDBM342442 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Cell-Free Assay |
|---|
| IC50 | 2263±n/a nM |
|---|
| Citation |  Brodney, MA; Butler, CR; Zhang, L; O Neill, BT N-(2-(2-amino-6-substituted-4,4a,5,6-tetrahydropyrano[3,4-d][1,3]OXAZIN-8a(8H)-yl)-thiazol-4-yl) amides US Patent US9771379 Publication Date 9/26/2017 Brodney, MA; Butler, CR; Zhang, L; O Neill, BT N-(2-(2-amino-6-substituted-4,4a,5,6-tetrahydropyrano[3,4-d][1,3]OXAZIN-8a(8H)-yl)-thiazol-4-yl) amides US Patent US9771379 Publication Date 9/26/2017 |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Beta-secretase 1 |
|---|
| Name: | Beta-secretase 1 |
|---|
| Synonyms: | ASP2 | Asp 2 | Aspartyl protease 2 | BACE | BACE1 | BACE1_HUMAN | Beta-secretase (BACE) | Beta-secretase 1 | Beta-secretase 1 (BACE 1) | Beta-secretase 1 (BACE-1) | Beta-secretase 1 (BACE1) | Beta-site APP cleaving enzyme 1 | Beta-site amyloid precursor protein cleaving enzyme 1 | KIAA1149 | Memapsin-2 | Membrane-associated aspartic protease 2 | beta-Secretase (BACE-1) | beta-Secretase (BACE1) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 55755.10 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P56817 |
|---|
| Residue: | 501 |
|---|
| Sequence: | MAQALPWLLLWMGAGVLPAHGTQHGIRLPLRSGLGGAPLGLRLPRETDEEPEEPGRRGSF
VEMVDNLRGKSGQGYYVEMTVGSPPQTLNILVDTGSSNFAVGAAPHPFLHRYYQRQLSST
YRDLRKGVYVPYTQGKWEGELGTDLVSIPHGPNVTVRANIAAITESDKFFINGSNWEGIL
GLAYAEIARPDDSLEPFFDSLVKQTHVPNLFSLQLCGAGFPLNQSEVLASVGGSMIIGGI
DHSLYTGSLWYTPIRREWYYEVIIVRVEINGQDLKMDCKEYNYDKSIVDSGTTNLRLPKK
VFEAAVKSIKAASSTEKFPDGFWLGEQLVCWQAGTTPWNIFPVISLYLMGEVTNQSFRIT
ILPQQYLRPVEDVATSQDDCYKFAISQSSTGTVMGAVIMEGFYVVFDRARKRIGFAVSAC
HVHDEFRTAAVEGPFVTLDMEDCGYNIPQTDESTLMTIAYVMAAICALFMLPLCLMVCQW
RCLRCLRQQHDDFADDISLLK
|
|
|
|---|
| BDBM342442 |
|---|
| n/a |
|---|
| Name | BDBM342442 |
|---|
| Synonyms: | N-(2-((4a'R,8a'R)-2'-amino- 4a',5'-dihydro-4'H-spiro [cyclobutane-1,6'-pyrano [3,4-d][1,3]oxazin]-8a'(8'H)- yl)thiazol-4-yl)-5- (difluoromethoxy)pyrazine- 2-carboxamide | US9771379, Example 17 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C19H20F2N6O4S |
|---|
| Mol. Mass. | 466.462 |
|---|
| SMILES | NC1=N[C@]2(COC3(CCC3)C[C@H]2CO1)c1nc(NC(=O)c2cnc(OC(F)F)cn2)cs1 |r,t:1| |
|---|
| Structure | 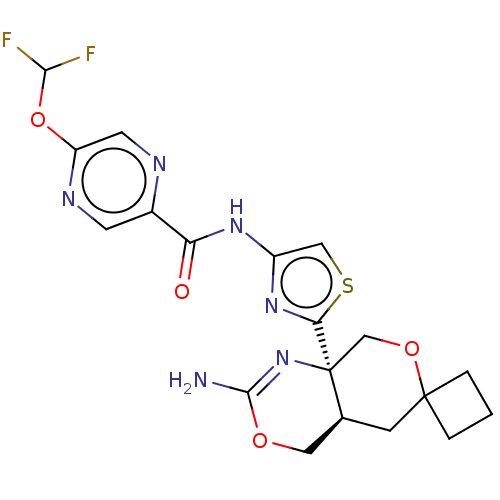
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Brodney, MA; Butler, CR; Zhang, L; O Neill, BT N-(2-(2-amino-6-substituted-4,4a,5,6-tetrahydropyrano[3,4-d][1,3]OXAZIN-8a(8H)-yl)-thiazol-4-yl) amides US Patent US9771379 Publication Date 9/26/2017
Brodney, MA; Butler, CR; Zhang, L; O Neill, BT N-(2-(2-amino-6-substituted-4,4a,5,6-tetrahydropyrano[3,4-d][1,3]OXAZIN-8a(8H)-yl)-thiazol-4-yl) amides US Patent US9771379 Publication Date 9/26/2017