| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Methionine aminopeptidase 2 |
|---|
| Ligand | BDBM17456 |
|---|
| Substrate/Competitor | BDBM17353 |
|---|
| Meas. Tech. | Enzyme Inhibition Assay |
|---|
| pH | 7.5±n/a |
|---|
| Temperature | 295.15±n/a K |
|---|
| Ki | >1000±n/a nM |
|---|
| Citation |  Kallander, LS; Lu, Q; Chen, W; Tomaszek, T; Yang, G; Tew, D; Meek, TD; Hofmann, GA; Schulz-Pritchard, CK; Smith, WW; Janson, CA; Ryan, MD; Zhang, GF; Johanson, KO; Kirkpatrick, RB; Ho, TF; Fisher, PW; Mattern, MR; Johnson, RK; Hansbury, MJ; Winkler, JD; Ward, KW; Veber, DF; Thompson, SK 4-Aryl-1,2,3-triazole: a novel template for a reversible methionine aminopeptidase 2 inhibitor, optimized to inhibit angiogenesis in vivo. J Med Chem48:5644-7 (2005) [PubMed] Article Kallander, LS; Lu, Q; Chen, W; Tomaszek, T; Yang, G; Tew, D; Meek, TD; Hofmann, GA; Schulz-Pritchard, CK; Smith, WW; Janson, CA; Ryan, MD; Zhang, GF; Johanson, KO; Kirkpatrick, RB; Ho, TF; Fisher, PW; Mattern, MR; Johnson, RK; Hansbury, MJ; Winkler, JD; Ward, KW; Veber, DF; Thompson, SK 4-Aryl-1,2,3-triazole: a novel template for a reversible methionine aminopeptidase 2 inhibitor, optimized to inhibit angiogenesis in vivo. J Med Chem48:5644-7 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Inhibition_Run data, Solution Info, Assay Method |
|---|
| |
| Methionine aminopeptidase 2 |
|---|
| Name: | Methionine aminopeptidase 2 |
|---|
| Synonyms: | Initiation factor 2-associated 67 kDa glycoprotein | MAP 2 | MAP2_HUMAN | METAP2 | MNPEP | MetAP 2 | Methionine aminopeptidase 2 (MetAP2) | Methionine aminopeptidases (HsMetAP2) | P67EIF2 | Peptidase M 2 | p67 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 52884.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P50579 |
|---|
| Residue: | 478 |
|---|
| Sequence: | MAGVEEVAASGSHLNGDLDPDDREEGAASTAEEAAKKKRRKKKKSKGPSAAGEQEPDKES
GASVDEVARQLERSALEDKERDEDDEDGDGDGDGATGKKKKKKKKKRGPKVQTDPPSVPI
CDLYPNGVFPKGQECEYPPTQDGRTAAWRTTSEEKKALDQASEEIWNDFREAAEAHRQVR
KYVMSWIKPGMTMIEICEKLEDCSRKLIKENGLNAGLAFPTGCSLNNCAAHYTPNAGDTT
VLQYDDICKIDFGTHISGRIIDCAFTVTFNPKYDTLLKAVKDATNTGIKCAGIDVRLCDV
GEAIQEVMESYEVEIDGKTYQVKPIRNLNGHSIGQYRIHAGKTVPIVKGGEATRMEEGEV
YAIETFGSTGKGVVHDDMECSHYMKNFDVGHVPIRLPRTKHLLNVINENFGTLAFCRRWL
DRLGESKYLMALKNLCDLGIVDPYPPLCDIKGSYTAQFEHTILLRPTCKEVVSRGDDY
|
|
|
|---|
| BDBM17456 |
|---|
| BDBM17353 |
|---|
| Name | BDBM17456 |
|---|
| Synonyms: | 1,2,3-triazole analogue, 12 | 5-(4-propylphenyl)-1H-1,2,3-triazole |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C11H13N3 |
|---|
| Mol. Mass. | 187.241 |
|---|
| SMILES | CCCc1ccc(cc1)-c1c[nH]nn1 |
|---|
| Structure | 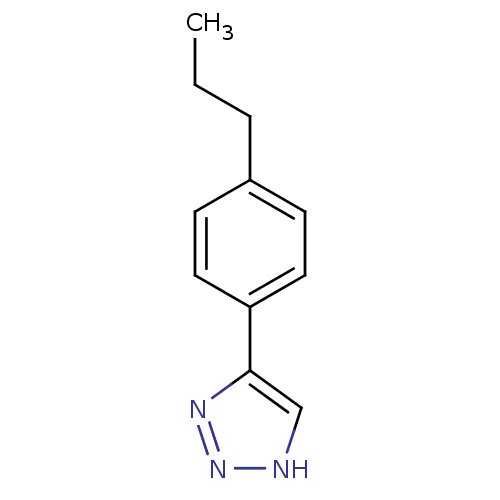
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Kallander, LS; Lu, Q; Chen, W; Tomaszek, T; Yang, G; Tew, D; Meek, TD; Hofmann, GA; Schulz-Pritchard, CK; Smith, WW; Janson, CA; Ryan, MD; Zhang, GF; Johanson, KO; Kirkpatrick, RB; Ho, TF; Fisher, PW; Mattern, MR; Johnson, RK; Hansbury, MJ; Winkler, JD; Ward, KW; Veber, DF; Thompson, SK 4-Aryl-1,2,3-triazole: a novel template for a reversible methionine aminopeptidase 2 inhibitor, optimized to inhibit angiogenesis in vivo. J Med Chem48:5644-7 (2005) [PubMed] Article
Kallander, LS; Lu, Q; Chen, W; Tomaszek, T; Yang, G; Tew, D; Meek, TD; Hofmann, GA; Schulz-Pritchard, CK; Smith, WW; Janson, CA; Ryan, MD; Zhang, GF; Johanson, KO; Kirkpatrick, RB; Ho, TF; Fisher, PW; Mattern, MR; Johnson, RK; Hansbury, MJ; Winkler, JD; Ward, KW; Veber, DF; Thompson, SK 4-Aryl-1,2,3-triazole: a novel template for a reversible methionine aminopeptidase 2 inhibitor, optimized to inhibit angiogenesis in vivo. J Med Chem48:5644-7 (2005) [PubMed] Article