| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Poly(ADP-ribose) glycohydrolase |
|---|
| Ligand | BDBM50466568 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1793403 (CHEMBL4265322) |
|---|
| EC50 | >70000±n/a nM |
|---|
| Citation |  Waszkowycz, B; Smith, KM; McGonagle, AE; Jordan, AM; Acton, B; Fairweather, EE; Griffiths, LA; Hamilton, NM; Hamilton, NS; Hitchin, JR; Hutton, CP; James, DI; Jones, CD; Jones, S; Mould, DP; Small, HF; Stowell, AIJ; Tucker, JA; Waddell, ID; Ogilvie, DJ Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides. J Med Chem61:10767-10792 (2018) [PubMed] Article Waszkowycz, B; Smith, KM; McGonagle, AE; Jordan, AM; Acton, B; Fairweather, EE; Griffiths, LA; Hamilton, NM; Hamilton, NS; Hitchin, JR; Hutton, CP; James, DI; Jones, CD; Jones, S; Mould, DP; Small, HF; Stowell, AIJ; Tucker, JA; Waddell, ID; Ogilvie, DJ Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides. J Med Chem61:10767-10792 (2018) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Poly(ADP-ribose) glycohydrolase |
|---|
| Name: | Poly(ADP-ribose) glycohydrolase |
|---|
| Synonyms: | PARG | PARG_HUMAN | Poly(ADP-ribose) glycohydrolase | poly(ADP-ribose) glycohydrolase (PARG) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 111107.13 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q86W56 |
|---|
| Residue: | 976 |
|---|
| Sequence: | MNAGPGCEPCTKRPRWGAATTSPAASDARSFPSRQRRVLDPKDAHVQFRVPPSSPACVPG
RAGQHRGSATSLVFKQKTITSWMDTKGIKTAESESLDSKENNNTRIESMMSSVQKDNFYQ
HNVEKLENVSQLSLDKSPTEKSTQYLNQHQTAAMCKWQNEGKHTEQLLESEPQTVTLVPE
QFSNANIDRSPQNDDHSDTDSEENRDNQQFLTTVKLANAKQTTEDEQAREAKSHQKCSKS
CDPGEDCASCQQDEIDVVPESPLSDVGSEDVGTGPKNDNKLTRQESCLGNSPPFEKESEP
ESPMDVDNSKNSCQDSEADEETSPGFDEQEDGSSSQTANKPSRFQARDADIEFRKRYSTK
GGEVRLHFQFEGGESRTGMNDLNAKLPGNISSLNVECRNSKQHGKKDSKITDHFMRLPKA
EDRRKEQWETKHQRTERKIPKYVPPHLSPDKKWLGTPIEEMRRMPRCGIRLPLLRPSANH
TVTIRVDLLRAGEVPKPFPTHYKDLWDNKHVKMPCSEQNLYPVEDENGERTAGSRWELIQ
TALLNKFTRPQNLKDAILKYNVAYSKKWDFTALIDFWDKVLEEAEAQHLYQSILPDMVKI
ALCLPNICTQPIPLLKQKMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDIN
FNRLFEGRSSRKPEKLKTLFCYFRRVTEKKPTGLVTFTRQSLEDFPEWERCEKPLTRLHV
TYEGTIEENGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIISRLFTEVLDHNEC
LIITGTEQYSEYTGYAETYRWSRSHEDGSERDDWQRRCTEIVAIDALHFRRYLDQFVPEK
MRRELNKAYCGFLRPGVSSENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERDVVYF
TFGDSELMRDIYSMHIFLTERKLTVGDVYKLLLRYYNEECRNCSTPGPDIKLYPFIYHAV
ESCAETADHSGQRTGT
|
|
|
|---|
| BDBM50466568 |
|---|
| n/a |
|---|
| Name | BDBM50466568 |
|---|
| Synonyms: | CHEMBL4288328 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H13Cl3N2O4S2 |
|---|
| Mol. Mass. | 527.828 |
|---|
| SMILES | OC(=O)CCN1C(=S)S\C(C1=O)=C1/C(=O)N(Cc2c(Cl)cccc2Cl)c2ccc(Cl)cc12 |
|---|
| Structure | 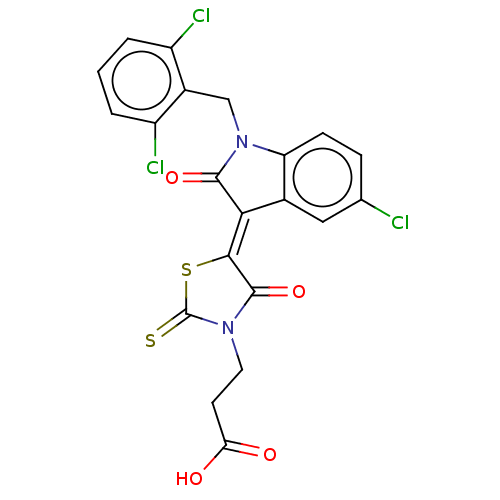
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Waszkowycz, B; Smith, KM; McGonagle, AE; Jordan, AM; Acton, B; Fairweather, EE; Griffiths, LA; Hamilton, NM; Hamilton, NS; Hitchin, JR; Hutton, CP; James, DI; Jones, CD; Jones, S; Mould, DP; Small, HF; Stowell, AIJ; Tucker, JA; Waddell, ID; Ogilvie, DJ Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides. J Med Chem61:10767-10792 (2018) [PubMed] Article
Waszkowycz, B; Smith, KM; McGonagle, AE; Jordan, AM; Acton, B; Fairweather, EE; Griffiths, LA; Hamilton, NM; Hamilton, NS; Hitchin, JR; Hutton, CP; James, DI; Jones, CD; Jones, S; Mould, DP; Small, HF; Stowell, AIJ; Tucker, JA; Waddell, ID; Ogilvie, DJ Cell-Active Small Molecule Inhibitors of the DNA-Damage Repair Enzyme Poly(ADP-ribose) Glycohydrolase (PARG): Discovery and Optimization of Orally Bioavailable Quinazolinedione Sulfonamides. J Med Chem61:10767-10792 (2018) [PubMed] Article