| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Prothrombin |
|---|
| Ligand | BDBM50093147 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_208334 |
|---|
| Ki | 492000±n/a nM |
|---|
| Citation |  Rice, KD; Gangloff, AR; Kuo, EY; Dener, JM; Wang, VR; Lum, R; Newcomb, WS; Havel, C; Putnam, D; Cregar, L; Wong, M; Warne, RL Dibasic inhibitors of human mast cell tryptase. Part 1: synthesis and optimization of a novel class of inhibitors. Bioorg Med Chem Lett10:2357-60 (2001) [PubMed] Rice, KD; Gangloff, AR; Kuo, EY; Dener, JM; Wang, VR; Lum, R; Newcomb, WS; Havel, C; Putnam, D; Cregar, L; Wong, M; Warne, RL Dibasic inhibitors of human mast cell tryptase. Part 1: synthesis and optimization of a novel class of inhibitors. Bioorg Med Chem Lett10:2357-60 (2001) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Prothrombin |
|---|
| Name: | Prothrombin |
|---|
| Synonyms: | Activation peptide fragment 1 | Activation peptide fragment 2 | Coagulation factor II | F2 | Prothrombin precursor | THRB_HUMAN | Thrombin heavy chain | Thrombin light chain |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 70029.57 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P00734 |
|---|
| Residue: | 622 |
|---|
| Sequence: | MAHVRGLQLPGCLALAALCSLVHSQHVFLAPQQARSLLQRVRRANTFLEEVRKGNLEREC
VEETCSYEEAFEALESSTATDVFWAKYTACETARTPRDKLAACLEGNCAEGLGTNYRGHV
NITRSGIECQLWRSRYPHKPEINSTTHPGADLQENFCRNPDSSTTGPWCYTTDPTVRRQE
CSIPVCGQDQVTVAMTPRSEGSSVNLSPPLEQCVPDRGQQYQGRLAVTTHGLPCLAWASA
QAKALSKHQDFNSAVQLVENFCRNPDGDEEGVWCYVAGKPGDFGYCDLNYCEEAVEEETG
DGLDEDSDRAIEGRTATSEYQTFFNPRTFGSGEADCGLRPLFEKKSLEDKTERELLESYI
DGRIVEGSDAEIGMSPWQVMLFRKSPQELLCGASLISDRWVLTAAHCLLYPPWDKNFTEN
DLLVRIGKHSRTRYERNIEKISMLEKIYIHPRYNWRENLDRDIALMKLKKPVAFSDYIHP
VCLPDRETAASLLQAGYKGRVTGWGNLKETWTANVGKGQPSVLQVVNLPIVERPVCKDST
RIRITDNMFCAGYKPDEGKRGDACEGDSGGPFVMKSPFNNRWYQMGIVSWGEGCDRDGKY
GFYTHVFRLKKWIQKVIDQFGE
|
|
|
|---|
| BDBM50093147 |
|---|
| n/a |
|---|
| Name | BDBM50093147 |
|---|
| Synonyms: | CHEMBL79545 | Pentanedioic acid 4-guanidino-benzylamide 4-{[4-(4-guanidino-benzylcarbamoyl)-butyrylamino]-methyl}-benzylamide |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C34H44N10O4 |
|---|
| Mol. Mass. | 656.7778 |
|---|
| SMILES | [#7]\[#6](-[#7])=[#7]/c1ccc(-[#6]-[#7]-[#6](=O)-[#6]-[#6]-[#6]-[#6](=O)-[#7]-[#6]-c2ccc(-[#6]-[#7]-[#6](=O)-[#6]-[#6]-[#6]-[#6](=O)-[#7]-[#6]-c3ccc(cc3)\[#7]=[#6](/[#7])-[#7])cc2)cc1 |
|---|
| Structure | 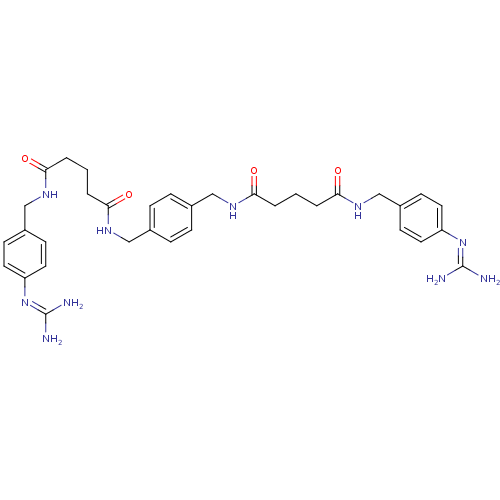
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Rice, KD; Gangloff, AR; Kuo, EY; Dener, JM; Wang, VR; Lum, R; Newcomb, WS; Havel, C; Putnam, D; Cregar, L; Wong, M; Warne, RL Dibasic inhibitors of human mast cell tryptase. Part 1: synthesis and optimization of a novel class of inhibitors. Bioorg Med Chem Lett10:2357-60 (2001) [PubMed]
Rice, KD; Gangloff, AR; Kuo, EY; Dener, JM; Wang, VR; Lum, R; Newcomb, WS; Havel, C; Putnam, D; Cregar, L; Wong, M; Warne, RL Dibasic inhibitors of human mast cell tryptase. Part 1: synthesis and optimization of a novel class of inhibitors. Bioorg Med Chem Lett10:2357-60 (2001) [PubMed]