| Reaction Details |
|---|
 | Report a problem with these data |
| Target | TGF-beta receptor type-1 |
|---|
| Ligand | BDBM50151352 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_306809 (CHEMBL830166) |
|---|
| IC50 | 380±n/a nM |
|---|
| Citation |  Gellibert, F; Woolven, J; Fouchet, MH; Mathews, N; Goodland, H; Lovegrove, V; Laroze, A; Nguyen, VL; Sautet, S; Wang, R; Janson, C; Smith, W; Krysa, G; Boullay, V; De Gouville, AC; Huet, S; Hartley, D Identification of 1,5-naphthyridine derivatives as a novel series of potent and selective TGF-beta type I receptor inhibitors. J Med Chem47:4494-506 (2004) [PubMed] Article Gellibert, F; Woolven, J; Fouchet, MH; Mathews, N; Goodland, H; Lovegrove, V; Laroze, A; Nguyen, VL; Sautet, S; Wang, R; Janson, C; Smith, W; Krysa, G; Boullay, V; De Gouville, AC; Huet, S; Hartley, D Identification of 1,5-naphthyridine derivatives as a novel series of potent and selective TGF-beta type I receptor inhibitors. J Med Chem47:4494-506 (2004) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| TGF-beta receptor type-1 |
|---|
| Name: | TGF-beta receptor type-1 |
|---|
| Synonyms: | ALK-5 | ALK5 | Activin A receptor type II-like protein kinase of 53kD | Activin receptor-like kinase 5 | SKR4 | Serine/threonine-protein kinase receptor R4 | TGF-beta receptor type I | TGF-beta type I receptor | TGFBR1 | TGFR-1 | TGFR1_HUMAN | TbetaR-I | Transforming growth factor-beta receptor type I |
|---|
| Type: | enzyme |
|---|
| Mol. Mass.: | 55968.24 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P36897 |
|---|
| Residue: | 503 |
|---|
| Sequence: | MEAAVAAPRPRLLLLVLAAAAAAAAALLPGATALQCFCHLCTKDNFTCVTDGLCFVSVTE
TTDKVIHNSMCIAEIDLIPRDRPFVCAPSSKTGSVTTTYCCNQDHCNKIELPTTVKSSPG
LGPVELAAVIAGPVCFVCISLMLMVYICHNRTVIHHRVPNEEDPSLDRPFISEGTTLKDL
IYDMTTSGSGSGLPLLVQRTIARTIVLQESIGKGRFGEVWRGKWRGEEVAVKIFSSREER
SWFREAEIYQTVMLRHENILGFIAADNKDNGTWTQLWLVSDYHEHGSLFDYLNRYTVTVE
GMIKLALSTASGLAHLHMEIVGTQGKPAIAHRDLKSKNILVKKNGTCCIADLGLAVRHDS
ATDTIDIAPNHRVGTKRYMAPEVLDDSINMKHFESFKRADIYAMGLVFWEIARRCSIGGI
HEDYQLPYYDLVPSDPSVEEMRKVVCEQKLRPNIPNRWQSCEALRVMAKIMRECWYANGA
ARLTALRIKKTLSQLSQQEGIKM
|
|
|
|---|
| BDBM50151352 |
|---|
| n/a |
|---|
| Name | BDBM50151352 |
|---|
| Synonyms: | 5-(6-Methyl-[1,5]naphthyridin-4-yl)-4-pyridin-2-yl-thiazol-2-ylamine | 5-(6-methyl-1,5-naphthyridin-4-yl)-4-(pyridin-2-yl)thiazol-2-amine | CHEMBL425177 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C17H13N5S |
|---|
| Mol. Mass. | 319.384 |
|---|
| SMILES | Cc1ccc2nccc(-c3sc(N)nc3-c3ccccn3)c2n1 |
|---|
| Structure | 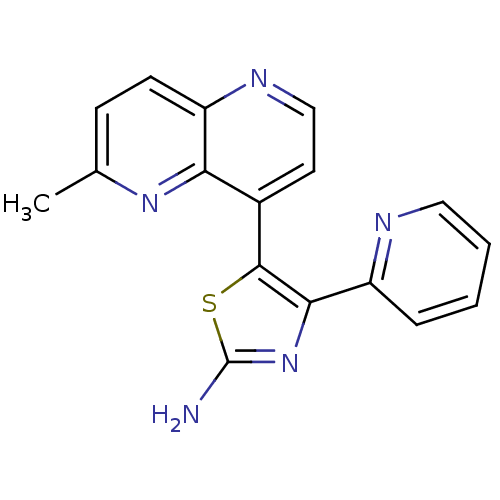
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Gellibert, F; Woolven, J; Fouchet, MH; Mathews, N; Goodland, H; Lovegrove, V; Laroze, A; Nguyen, VL; Sautet, S; Wang, R; Janson, C; Smith, W; Krysa, G; Boullay, V; De Gouville, AC; Huet, S; Hartley, D Identification of 1,5-naphthyridine derivatives as a novel series of potent and selective TGF-beta type I receptor inhibitors. J Med Chem47:4494-506 (2004) [PubMed] Article
Gellibert, F; Woolven, J; Fouchet, MH; Mathews, N; Goodland, H; Lovegrove, V; Laroze, A; Nguyen, VL; Sautet, S; Wang, R; Janson, C; Smith, W; Krysa, G; Boullay, V; De Gouville, AC; Huet, S; Hartley, D Identification of 1,5-naphthyridine derivatives as a novel series of potent and selective TGF-beta type I receptor inhibitors. J Med Chem47:4494-506 (2004) [PubMed] Article