

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Glutathione S-transferase P/Lysine-specific histone demethylase 1A [158-852] | ||
| Ligand | BDBM254554 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | Biological Assay | ||
| pH | 7.4±n/a | ||
| Temperature | 310.15±n/a K | ||
| Ki | 550±n/a nM | ||
| Comments | extracted | ||
| Citation |  Ortega Muñoz, A; Fyfe, MC; Martinell Pedemonte, M; Estiarte Martínez, M; Valls Vidal, N; Kurz, G; Castro Palomino Laria, JC (Hetero)aryl cyclopropylamine compounds as LSD1 inhibitors US Patent US9469597 Publication Date 10/18/2016 Ortega Muñoz, A; Fyfe, MC; Martinell Pedemonte, M; Estiarte Martínez, M; Valls Vidal, N; Kurz, G; Castro Palomino Laria, JC (Hetero)aryl cyclopropylamine compounds as LSD1 inhibitors US Patent US9469597 Publication Date 10/18/2016 | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Glutathione S-transferase P/Lysine-specific histone demethylase 1A [158-852] | |||
| Name: | Glutathione S-transferase P/Lysine-specific histone demethylase 1A [158-852] | ||
| Synonyms: | GST-Lysine Specific Demethylase-1 (LSD1) | ||
| Type: | Enzyme | ||
| Mol. Mass.: | n/a | ||
| Description: | n/a | ||
| Components: | This complex has 2 components. | ||
| Component 1 | |||
| Name: | Glutathione S-transferase P | ||
| Synonyms: | FAEES3 | GST class-pi | GST3 | GSTP1 | GSTP1-1 | GSTP1_HUMAN | Glutathione S-transferase | Glutathione S-transferase (GST) | Glutathione S-transferase P | Glutathione S-transferase Pi | Glutathione transferase (GST) | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 23353.53 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P09211 | ||
| Residue: | 210 | ||
| Sequence: |
| ||
| Component 2 | |||
| Name: | Lysine-specific histone demethylase 1A [158-852] | ||
| Synonyms: | AOF2 | KDM1 | KDM1A | KDM1A_HUMAN | KIAA0601 | LSD1 | Lysine Specific Demethylase-1 (LSD1) | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 77140.81 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | aa 158-852 | ||
| Residue: | 695 | ||
| Sequence: |
| ||
| BDBM254554 | |||
| n/a | |||
| Name | BDBM254554 | ||
| Synonyms: | US10214477, Example 13 | US9469597, 13 | US9670136, 13 N-(4-(((trans)-2-(6-(3-(trifluoromethyl)phenyl)pyridin-3-yl)cyclopropyl)amino)cyclohexyl)acetamide | ||
| Type | Small organic molecule | ||
| Emp. Form. | C23H26F3N3O | ||
| Mol. Mass. | 417.4672 | ||
| SMILES | CC(=O)NC1CCC(CC1)N[C@H]1C[C@@H]1c1ccc(nc1)-c1cccc(c1)C(F)(F)F |r,wU:13.15,wD:11.11,(9.54,5.39,;8.21,4.62,;8.21,3.08,;6.87,5.39,;5.54,4.62,;5.54,3.08,;4.21,2.31,;2.87,3.08,;2.87,4.62,;4.21,5.39,;1.54,2.31,;.21,3.08,;-.56,4.41,;-1.33,3.08,;-2.67,2.31,;-4,3.08,;-5.33,2.31,;-5.33,.77,;-4,,;-2.67,.77,;-6.67,,;-8,.77,;-9.34,,;-9.34,-1.54,;-8,-2.37,;-6.67,-1.54,;-8,-3.91,;-8,-5.45,;-9.54,-3.91,;-6.46,-3.91,)| | ||
| Structure | 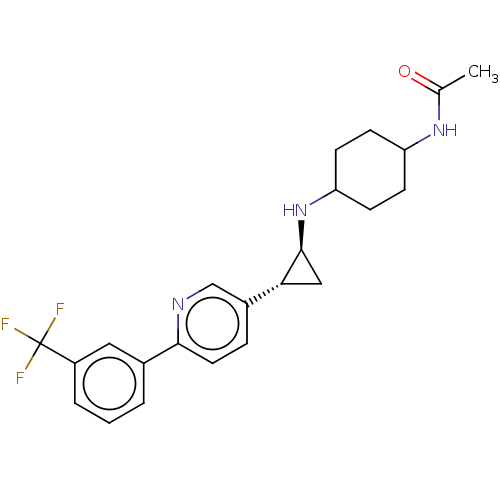
| ||