| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor delta |
|---|
| Ligand | BDBM50227744 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_452359 (CHEMBL902598) |
|---|
| IC50 | 3±n/a nM |
|---|
| Citation |  Shi, Q; Canada, EJ; Xu, Y; Warshawsky, AM; Etgen, GJ; Broderick, CL; Clutinger, CK; Irwin, LA; Laurila, ME; Montrose-Rafizadeh, C; Oldham, BA; Wang, M; Winneroski, LL; Xie, C; York, JS; Yumibe, NP; Zink, RW; Mantlo, N Design and synthesis of novel and potent amide linked PPARgamma/delta dual agonists. Bioorg Med Chem Lett17:6744-9 (2008) [PubMed] Article Shi, Q; Canada, EJ; Xu, Y; Warshawsky, AM; Etgen, GJ; Broderick, CL; Clutinger, CK; Irwin, LA; Laurila, ME; Montrose-Rafizadeh, C; Oldham, BA; Wang, M; Winneroski, LL; Xie, C; York, JS; Yumibe, NP; Zink, RW; Mantlo, N Design and synthesis of novel and potent amide linked PPARgamma/delta dual agonists. Bioorg Med Chem Lett17:6744-9 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor delta |
|---|
| Name: | Peroxisome proliferator-activated receptor delta |
|---|
| Synonyms: | NR1C2 | NUC1 | NUCI | Nuclear hormone receptor 1 | Nuclear receptor subfamily 1 group C member 2 | PPAR delta | PPAR-beta | PPARB | PPARD | PPARD_HUMAN | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor beta | Peroxisome proliferator-activated receptor delta |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 49910.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q03181 |
|---|
| Residue: | 441 |
|---|
| Sequence: | MEQPQEEAPEVREEEEKEEVAEAEGAPELNGGPQHALPSSSYTDLSRSSSPPSLLDQLQM
GCDGASCGSLNMECRVCGDKASGFHYGVHACEGCKGFFRRTIRMKLEYEKCERSCKIQKK
NRNKCQYCRFQKCLALGMSHNAIRFGRMPEAEKRKLVAGLTANEGSQYNPQVADLKAFSK
HIYNAYLKNFNMTKKKARSILTGKASHTAPFVIHDIETLWQAEKGLVWKQLVNGLPPYKE
ISVHVFYRCQCTTVETVRELTEFAKSIPSFSSLFLNDQVTLLKYGVHEAIFAMLASIVNK
DGLLVANGSGFVTREFLRSLRKPFSDIIEPKFEFAVKFNALELDDSDLALFIAAIILCGD
RPGLMNVPRVEAIQDTILRALEFHLQANHPDAQYLFPKLLQKMADLRQLVTEHAQMMQRI
KKTETETSLHPLLQEIYKDMY
|
|
|
|---|
| BDBM50227744 |
|---|
| n/a |
|---|
| Name | BDBM50227744 |
|---|
| Synonyms: | 3-(4-(3-((5-chloro-1,3-dimethyl-1H-indole-2-carboxamido)methyl)phenoxy)-2-methylphenyl)propanoic acid | CHEMBL238152 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C28H27ClN2O4 |
|---|
| Mol. Mass. | 490.978 |
|---|
| SMILES | Cc1c(C(=O)NCc2cccc(Oc3ccc(CCC(O)=O)c(C)c3)c2)n(C)c2ccc(Cl)cc12 |
|---|
| Structure | 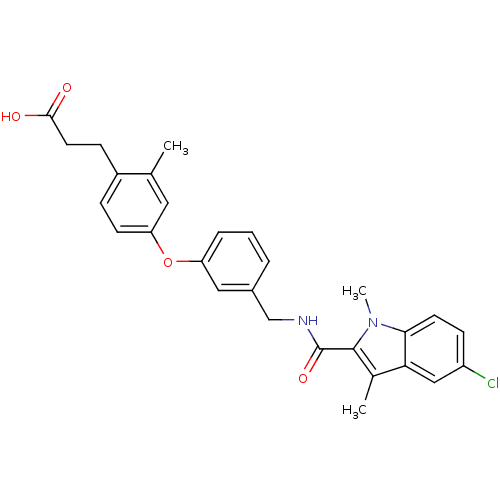
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Shi, Q; Canada, EJ; Xu, Y; Warshawsky, AM; Etgen, GJ; Broderick, CL; Clutinger, CK; Irwin, LA; Laurila, ME; Montrose-Rafizadeh, C; Oldham, BA; Wang, M; Winneroski, LL; Xie, C; York, JS; Yumibe, NP; Zink, RW; Mantlo, N Design and synthesis of novel and potent amide linked PPARgamma/delta dual agonists. Bioorg Med Chem Lett17:6744-9 (2008) [PubMed] Article
Shi, Q; Canada, EJ; Xu, Y; Warshawsky, AM; Etgen, GJ; Broderick, CL; Clutinger, CK; Irwin, LA; Laurila, ME; Montrose-Rafizadeh, C; Oldham, BA; Wang, M; Winneroski, LL; Xie, C; York, JS; Yumibe, NP; Zink, RW; Mantlo, N Design and synthesis of novel and potent amide linked PPARgamma/delta dual agonists. Bioorg Med Chem Lett17:6744-9 (2008) [PubMed] Article