| Reaction Details |
|---|
 | Report a problem with these data |
| Target | 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase |
|---|
| Ligand | BDBM36495 |
|---|
| Substrate/Competitor | BDBM22111 |
|---|
| Meas. Tech. | Enzyme Activity Assay |
|---|
| pH | 7.5±0 |
|---|
| Temperature | 298.15±0 K |
|---|
| Kd | 0.012±0.0 nM |
|---|
| Citation |  Gutierrez, JA; Luo, M; Singh, V; Li, L; Brown, RL; Norris, GE; Evans, GB; Furneaux, RH; Tyler, PC; Painter, GF; Lenz, DH; Schramm, VL Picomolar inhibitors as transition-state probes of 5'-methylthioadenosine nucleosidases. ACS Chem Biol2:725-34 (2007) [PubMed] Article Gutierrez, JA; Luo, M; Singh, V; Li, L; Brown, RL; Norris, GE; Evans, GB; Furneaux, RH; Tyler, PC; Painter, GF; Lenz, DH; Schramm, VL Picomolar inhibitors as transition-state probes of 5'-methylthioadenosine nucleosidases. ACS Chem Biol2:725-34 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase |
|---|
| Name: | 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase |
|---|
| Synonyms: | 5 -methylthioadenosine nucleosidase | 5'-methylthioadenosine nucleosidase | 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase | 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase (MTAN) | MTA/SAH nucleosidase | MTAN | MTNN_ECOLI | Methylthioadenosine Nucleosidase(MTAN) | P46 | S-adenosylhomocysteine nucleosidase | mtn | mtnN | pfs | yadA |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 24347.14 |
|---|
| Organism: | Escherichia coli (strain K12) |
|---|
| Description: | P0AF12 |
|---|
| Residue: | 232 |
|---|
| Sequence: | MKIGIIGAMEEEVTLLRDKIENRQTISLGGCEIYTGQLNGTEVALLKSGIGKVAAALGAT
LLLEHCKPDVIINTGSAGGLAPTLKVGDIVVSDEARYHDADVTAFGYEYGQLPGCPAGFK
ADDKLIAAAEACIAELNLNAVRGLIVSGDAFINGSVGLAKIRHNFPQAIAVEMEATAIAH
VCHNFNVPFVVVRAISDVADQQSHLSFDEFLAVAAKQSSLMVESLVQKLAHG
|
|
|
|---|
| BDBM36495 |
|---|
| BDBM22111 |
|---|
| Name | BDBM36495 |
|---|
| Synonyms: | CHEMBL554642 | ImmA-Bn |
|---|
| Type | Small organic moelcule |
|---|
| Emp. Form. | C18H21N5O2S |
|---|
| Mol. Mass. | 371.457 |
|---|
| SMILES | Nc1ncnc2c(c[nH]c12)C1NC(CSCc2ccccc2)C(O)C1O |
|---|
| Structure | 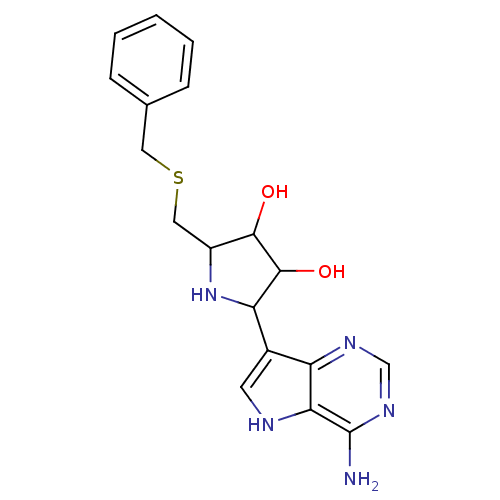
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Gutierrez, JA; Luo, M; Singh, V; Li, L; Brown, RL; Norris, GE; Evans, GB; Furneaux, RH; Tyler, PC; Painter, GF; Lenz, DH; Schramm, VL Picomolar inhibitors as transition-state probes of 5'-methylthioadenosine nucleosidases. ACS Chem Biol2:725-34 (2007) [PubMed] Article
Gutierrez, JA; Luo, M; Singh, V; Li, L; Brown, RL; Norris, GE; Evans, GB; Furneaux, RH; Tyler, PC; Painter, GF; Lenz, DH; Schramm, VL Picomolar inhibitors as transition-state probes of 5'-methylthioadenosine nucleosidases. ACS Chem Biol2:725-34 (2007) [PubMed] Article