| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Beta-secretase 1 |
|---|
| Ligand | BDBM50250536 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1681840 (CHEMBL4032117) |
|---|
| IC50 | 90±n/a nM |
|---|
| Citation |  Bueno, AB; Agejas, J; Broughton, H; Dally, R; Durham, TB; Espinosa, JF; Gonz�lez, R; Hahn, PJ; Marcos, A; Rodr�guez, R; Sanz, G; Soriano, JF; Timm, D; Vidal, P; Yang, HC; McCarthy, JR Optimization of Hydroxyethylamine Transition State Isosteres as Aspartic Protease Inhibitors by Exploiting Conformational Preferences. J Med Chem60:9807-9820 (2017) [PubMed] Article Bueno, AB; Agejas, J; Broughton, H; Dally, R; Durham, TB; Espinosa, JF; Gonz�lez, R; Hahn, PJ; Marcos, A; Rodr�guez, R; Sanz, G; Soriano, JF; Timm, D; Vidal, P; Yang, HC; McCarthy, JR Optimization of Hydroxyethylamine Transition State Isosteres as Aspartic Protease Inhibitors by Exploiting Conformational Preferences. J Med Chem60:9807-9820 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Beta-secretase 1 |
|---|
| Name: | Beta-secretase 1 |
|---|
| Synonyms: | ASP2 | Asp 2 | Aspartyl protease 2 | BACE | BACE1 | BACE1_HUMAN | Beta-secretase (BACE) | Beta-secretase 1 | Beta-secretase 1 (BACE 1) | Beta-secretase 1 (BACE-1) | Beta-secretase 1 (BACE1) | Beta-site APP cleaving enzyme 1 | Beta-site amyloid precursor protein cleaving enzyme 1 | KIAA1149 | Memapsin-2 | Membrane-associated aspartic protease 2 | beta-Secretase (BACE-1) | beta-Secretase (BACE1) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 55755.10 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P56817 |
|---|
| Residue: | 501 |
|---|
| Sequence: | MAQALPWLLLWMGAGVLPAHGTQHGIRLPLRSGLGGAPLGLRLPRETDEEPEEPGRRGSF
VEMVDNLRGKSGQGYYVEMTVGSPPQTLNILVDTGSSNFAVGAAPHPFLHRYYQRQLSST
YRDLRKGVYVPYTQGKWEGELGTDLVSIPHGPNVTVRANIAAITESDKFFINGSNWEGIL
GLAYAEIARPDDSLEPFFDSLVKQTHVPNLFSLQLCGAGFPLNQSEVLASVGGSMIIGGI
DHSLYTGSLWYTPIRREWYYEVIIVRVEINGQDLKMDCKEYNYDKSIVDSGTTNLRLPKK
VFEAAVKSIKAASSTEKFPDGFWLGEQLVCWQAGTTPWNIFPVISLYLMGEVTNQSFRIT
ILPQQYLRPVEDVATSQDDCYKFAISQSSTGTVMGAVIMEGFYVVFDRARKRIGFAVSAC
HVHDEFRTAAVEGPFVTLDMEDCGYNIPQTDESTLMTIAYVMAAICALFMLPLCLMVCQW
RCLRCLRQQHDDFADDISLLK
|
|
|
|---|
| BDBM50250536 |
|---|
| n/a |
|---|
| Name | BDBM50250536 |
|---|
| Synonyms: | CHEMBL4077530 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H33ClF2N2O4 |
|---|
| Mol. Mass. | 450.948 |
|---|
| SMILES | Cl.[H][C@@]1(CO[C@@H](OCC(C)(C)C)[C@H](C)N1)[C@@H](O)[C@H](Cc1cc(F)cc(F)c1)NC(C)=O |r| |
|---|
| Structure | 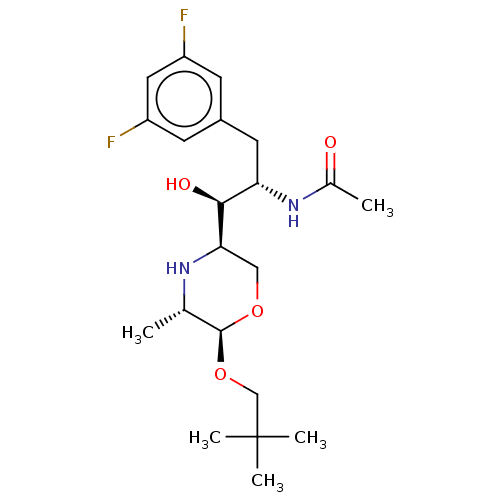
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Bueno, AB; Agejas, J; Broughton, H; Dally, R; Durham, TB; Espinosa, JF; Gonz�lez, R; Hahn, PJ; Marcos, A; Rodr�guez, R; Sanz, G; Soriano, JF; Timm, D; Vidal, P; Yang, HC; McCarthy, JR Optimization of Hydroxyethylamine Transition State Isosteres as Aspartic Protease Inhibitors by Exploiting Conformational Preferences. J Med Chem60:9807-9820 (2017) [PubMed] Article
Bueno, AB; Agejas, J; Broughton, H; Dally, R; Durham, TB; Espinosa, JF; Gonz�lez, R; Hahn, PJ; Marcos, A; Rodr�guez, R; Sanz, G; Soriano, JF; Timm, D; Vidal, P; Yang, HC; McCarthy, JR Optimization of Hydroxyethylamine Transition State Isosteres as Aspartic Protease Inhibitors by Exploiting Conformational Preferences. J Med Chem60:9807-9820 (2017) [PubMed] Article