| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Neuronal acetylcholine receptor subunit alpha-7 |
|---|
| Ligand | BDBM50451604 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1745955 (CHEMBL4180465) |
|---|
| EC50 | 32±n/a nM |
|---|
| Citation |  Cook, J; Zusi, FC; Hill, MD; Fang, H; Pearce, B; Park, H; Gallagher, L; McDonald, IM; Bristow, L; Macor, JE; Olson, RE Design and synthesis of a novel series of (1'S,2R,4'S)-3H-4'-azaspiro[benzo[4,5]imidazo[2,1-b]oxazole-2,2'-bicyclo[2.2.2]octanes] with high affinity for the ?7 neuronal nicotinic receptor. Bioorg Med Chem Lett27:5002-5005 (2017) [PubMed] Article Cook, J; Zusi, FC; Hill, MD; Fang, H; Pearce, B; Park, H; Gallagher, L; McDonald, IM; Bristow, L; Macor, JE; Olson, RE Design and synthesis of a novel series of (1'S,2R,4'S)-3H-4'-azaspiro[benzo[4,5]imidazo[2,1-b]oxazole-2,2'-bicyclo[2.2.2]octanes] with high affinity for the ?7 neuronal nicotinic receptor. Bioorg Med Chem Lett27:5002-5005 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Neuronal acetylcholine receptor subunit alpha-7 |
|---|
| Name: | Neuronal acetylcholine receptor subunit alpha-7 |
|---|
| Synonyms: | ACHA7_RAT | Acra7 | Cholinergic, Nicotinic Alpha7 | Cholinergic, Nicotinic Alpha7/5-HT3 | Chrna7 | Neuronal acetylcholine receptor | Neuronal acetylcholine receptor (alpha7 nAChR) | Neuronal acetylcholine receptor subunit alpha 7 | Neuronal acetylcholine receptor subunit alpha-7 | Neuronal acetylcholine receptor subunit alpha-7 (nAChR alpha7) | Neuronal acetylcholine receptor subunit alpha-7 (nAChR) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 56502.44 |
|---|
| Organism: | Rattus norvegicus (Rat) |
|---|
| Description: | Q05941 |
|---|
| Residue: | 502 |
|---|
| Sequence: | MCGGRGGIWLALAAALLHVSLQGEFQRRLYKELVKNYNPLERPVANDSQPLTVYFSLSLL
QIMDVDEKNQVLTTNIWLQMSWTDHYLQWNMSEYPGVKNVRFPDGQIWKPDILLYNSADE
RFDATFHTNVLVNASGHCQYLPPGIFKSSCYIDVRWFPFDVQQCKLKFGSWSYGGWSLDL
QMQEADISSYIPNGEWDLMGIPGKRNEKFYECCKEPYPDVTYTVTMRRRTLYYGLNLLIP
CVLISALALLVFLLPADSGEKISLGITVLLSLTVFMLLVAEIMPATSDSVPLIAQYFAST
MIIVGLSVVVTVIVLRYHHHDPDGGKMPKWTRIILLNWCAWFLRMKRPGEDKVRPACQHK
PRRCSLASVELSAGAGPPTSNGNLLYIGFRGLEGMHCAPTPDSGVVCGRLACSPTHDEHL
MHGAHPSDGDPDLAKILEEVRYIANRFRCQDESEVICSEWKFAACVVDRLCLMAFSVFTI
ICTIGILMSAPNFVEAVSKDFA
|
|
|
|---|
| BDBM50451604 |
|---|
| n/a |
|---|
| Name | BDBM50451604 |
|---|
| Synonyms: | CHEMBL4204677 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C11H13Br2N3O |
|---|
| Mol. Mass. | 363.048 |
|---|
| SMILES | Brc1nc2OC3(Cn2c1Br)CN1CCC3CC1 |THB:4:5:13.12:16.15,(20.16,-14,;18.86,-14.88,;17.41,-14.34,;16.47,-15.56,;14.99,-15.99,;14.94,-17.53,;16.38,-18.06,;17.33,-16.85,;18.81,-16.41,;20.06,-17.37,;15.56,-18.89,;14.01,-18.57,;12.59,-19.61,;12.06,-18.23,;13.42,-17.22,;13.1,-15.53,;13.83,-16.68,)| |
|---|
| Structure | 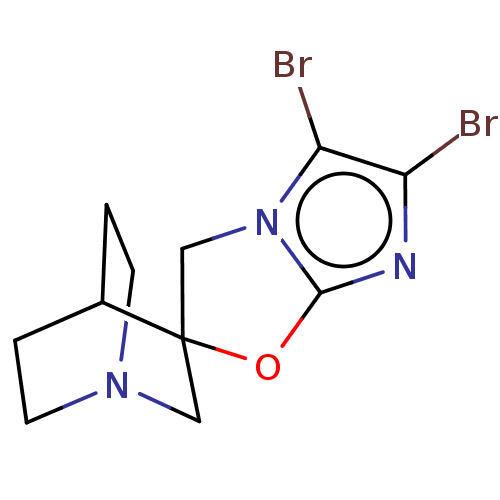
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Cook, J; Zusi, FC; Hill, MD; Fang, H; Pearce, B; Park, H; Gallagher, L; McDonald, IM; Bristow, L; Macor, JE; Olson, RE Design and synthesis of a novel series of (1'S,2R,4'S)-3H-4'-azaspiro[benzo[4,5]imidazo[2,1-b]oxazole-2,2'-bicyclo[2.2.2]octanes] with high affinity for the ?7 neuronal nicotinic receptor. Bioorg Med Chem Lett27:5002-5005 (2017) [PubMed] Article
Cook, J; Zusi, FC; Hill, MD; Fang, H; Pearce, B; Park, H; Gallagher, L; McDonald, IM; Bristow, L; Macor, JE; Olson, RE Design and synthesis of a novel series of (1'S,2R,4'S)-3H-4'-azaspiro[benzo[4,5]imidazo[2,1-b]oxazole-2,2'-bicyclo[2.2.2]octanes] with high affinity for the ?7 neuronal nicotinic receptor. Bioorg Med Chem Lett27:5002-5005 (2017) [PubMed] Article