| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Urokinase-type plasminogen activator |
|---|
| Ligand | BDBM50138668 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_213154 (CHEMBL815981) |
|---|
| Ki | 35.5±n/a nM |
|---|
| Citation |  Wendt, MD; Rockway, TW; Geyer, A; McClellan, W; Weitzberg, M; Zhao, X; Mantei, R; Nienaber, VL; Stewart, K; Klinghofer, V; Giranda, VL Identification of novel binding interactions in the development of potent, selective 2-naphthamidine inhibitors of urokinase. Synthesis, structural analysis, and SAR of N-phenyl amide 6-substitution. J Med Chem47:303-24 (2004) [PubMed] Article Wendt, MD; Rockway, TW; Geyer, A; McClellan, W; Weitzberg, M; Zhao, X; Mantei, R; Nienaber, VL; Stewart, K; Klinghofer, V; Giranda, VL Identification of novel binding interactions in the development of potent, selective 2-naphthamidine inhibitors of urokinase. Synthesis, structural analysis, and SAR of N-phenyl amide 6-substitution. J Med Chem47:303-24 (2004) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Urokinase-type plasminogen activator |
|---|
| Name: | Urokinase-type plasminogen activator |
|---|
| Synonyms: | 3.4.21.73 | PLAU | U-plasminogen activator | UROK_HUMAN | Urokinase | Urokinase-type plasminogen activator (uPA) | Urokinase-type plasminogen activator chain B | Urokinase-type plasminogen activator long chain A | Urokinase-type plasminogen activator short chain A | Urokinase-type plasminogen activator/surface receptor | uPA |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 48528.62 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P00749 |
|---|
| Residue: | 431 |
|---|
| Sequence: | MRALLARLLLCVLVVSDSKGSNELHQVPSNCDCLNGGTCVSNKYFSNIHWCNCPKKFGGQ
HCEIDKSKTCYEGNGHFYRGKASTDTMGRPCLPWNSATVLQQTYHAHRSDALQLGLGKHN
YCRNPDNRRRPWCYVQVGLKLLVQECMVHDCADGKKPSSPPEELKFQCGQKTLRPRFKII
GGEFTTIENQPWFAAIYRRHRGGSVTYVCGGSLISPCWVISATHCFIDYPKKEDYIVYLG
RSRLNSNTQGEMKFEVENLILHKDYSADTLAHHNDIALLKIRSKEGRCAQPSRTIQTICL
PSMYNDPQFGTSCEITGFGKENSTDYLYPEQLKMTVVKLISHRECQQPHYYGSEVTTKML
CAADPQWKTDSCQGDSGGPLVCSLQGRMTLTGIVSWGRGCALKDKPGVYTRVSHFLPWIR
SHTKEENGLAL
|
|
|
|---|
| BDBM50138668 |
|---|
| n/a |
|---|
| Name | BDBM50138668 |
|---|
| Synonyms: | 6-Carbamimidoyl-naphthalene-2-carboxylic acid (1-cyclopentyl-3,4-dihydro-isoquinolin-7-yl)-amide | CHEMBL323387 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H26N4O |
|---|
| Mol. Mass. | 410.5108 |
|---|
| SMILES | NC(=N)c1ccc2cc(ccc2c1)C(=O)Nc1ccc2CCN=C(C3CCCC3)c2c1 |t:24| |
|---|
| Structure | 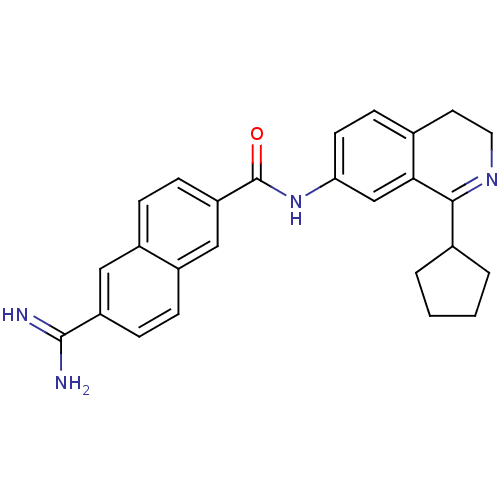
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Wendt, MD; Rockway, TW; Geyer, A; McClellan, W; Weitzberg, M; Zhao, X; Mantei, R; Nienaber, VL; Stewart, K; Klinghofer, V; Giranda, VL Identification of novel binding interactions in the development of potent, selective 2-naphthamidine inhibitors of urokinase. Synthesis, structural analysis, and SAR of N-phenyl amide 6-substitution. J Med Chem47:303-24 (2004) [PubMed] Article
Wendt, MD; Rockway, TW; Geyer, A; McClellan, W; Weitzberg, M; Zhao, X; Mantei, R; Nienaber, VL; Stewart, K; Klinghofer, V; Giranda, VL Identification of novel binding interactions in the development of potent, selective 2-naphthamidine inhibitors of urokinase. Synthesis, structural analysis, and SAR of N-phenyl amide 6-substitution. J Med Chem47:303-24 (2004) [PubMed] Article