| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor delta |
|---|
| Ligand | BDBM50171993 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_306113 |
|---|
| IC50 | >15000±n/a nM |
|---|
| Citation |  Shi, GQ; Dropinski, JF; Zhang, Y; Santini, C; Sahoo, SP; Berger, JP; Macnaul, KL; Zhou, G; Agrawal, A; Alvaro, R; Cai, TQ; Hernandez, M; Wright, SD; Moller, DE; Heck, JV; Meinke, PT Novel 2,3-dihydrobenzofuran-2-carboxylic acids: highly potent and subtype-selective PPARalpha agonists with potent hypolipidemic activity. J Med Chem48:5589-99 (2005) [PubMed] Article Shi, GQ; Dropinski, JF; Zhang, Y; Santini, C; Sahoo, SP; Berger, JP; Macnaul, KL; Zhou, G; Agrawal, A; Alvaro, R; Cai, TQ; Hernandez, M; Wright, SD; Moller, DE; Heck, JV; Meinke, PT Novel 2,3-dihydrobenzofuran-2-carboxylic acids: highly potent and subtype-selective PPARalpha agonists with potent hypolipidemic activity. J Med Chem48:5589-99 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor delta |
|---|
| Name: | Peroxisome proliferator-activated receptor delta |
|---|
| Synonyms: | NR1C2 | NUC1 | NUCI | Nuclear hormone receptor 1 | Nuclear receptor subfamily 1 group C member 2 | PPAR delta | PPAR-beta | PPARB | PPARD | PPARD_HUMAN | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor beta | Peroxisome proliferator-activated receptor delta |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 49910.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q03181 |
|---|
| Residue: | 441 |
|---|
| Sequence: | MEQPQEEAPEVREEEEKEEVAEAEGAPELNGGPQHALPSSSYTDLSRSSSPPSLLDQLQM
GCDGASCGSLNMECRVCGDKASGFHYGVHACEGCKGFFRRTIRMKLEYEKCERSCKIQKK
NRNKCQYCRFQKCLALGMSHNAIRFGRMPEAEKRKLVAGLTANEGSQYNPQVADLKAFSK
HIYNAYLKNFNMTKKKARSILTGKASHTAPFVIHDIETLWQAEKGLVWKQLVNGLPPYKE
ISVHVFYRCQCTTVETVRELTEFAKSIPSFSSLFLNDQVTLLKYGVHEAIFAMLASIVNK
DGLLVANGSGFVTREFLRSLRKPFSDIIEPKFEFAVKFNALELDDSDLALFIAAIILCGD
RPGLMNVPRVEAIQDTILRALEFHLQANHPDAQYLFPKLLQKMADLRQLVTEHAQMMQRI
KKTETETSLHPLLQEIYKDMY
|
|
|
|---|
| BDBM50171993 |
|---|
| n/a |
|---|
| Name | BDBM50171993 |
|---|
| Synonyms: | 5-[3-(2-Chloro-4-trifluoromethylsulfanyl-phenoxy)-propoxy]-2-ethyl-2,3-dihydro-benzofuran-2-carboxylic acid | CHEMBL425725 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H20ClF3O5S |
|---|
| Mol. Mass. | 476.894 |
|---|
| SMILES | CCC1(Cc2cc(OCCCOc3ccc(SC(F)(F)F)cc3Cl)ccc2O1)C(O)=O |
|---|
| Structure | 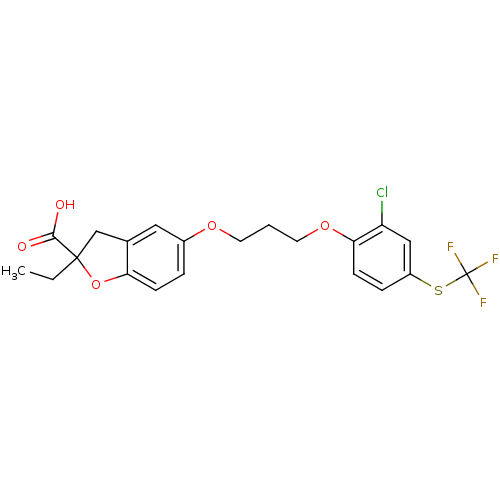
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Shi, GQ; Dropinski, JF; Zhang, Y; Santini, C; Sahoo, SP; Berger, JP; Macnaul, KL; Zhou, G; Agrawal, A; Alvaro, R; Cai, TQ; Hernandez, M; Wright, SD; Moller, DE; Heck, JV; Meinke, PT Novel 2,3-dihydrobenzofuran-2-carboxylic acids: highly potent and subtype-selective PPARalpha agonists with potent hypolipidemic activity. J Med Chem48:5589-99 (2005) [PubMed] Article
Shi, GQ; Dropinski, JF; Zhang, Y; Santini, C; Sahoo, SP; Berger, JP; Macnaul, KL; Zhou, G; Agrawal, A; Alvaro, R; Cai, TQ; Hernandez, M; Wright, SD; Moller, DE; Heck, JV; Meinke, PT Novel 2,3-dihydrobenzofuran-2-carboxylic acids: highly potent and subtype-selective PPARalpha agonists with potent hypolipidemic activity. J Med Chem48:5589-99 (2005) [PubMed] Article