| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Nociceptin receptor |
|---|
| Ligand | BDBM50181392 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_331367 (CHEMBL865854) |
|---|
| EC50 | >1000±n/a nM |
|---|
| Citation |  Goto, Y; Arai-Otsuki, S; Tachibana, Y; Ichikawa, D; Ozaki, S; Takahashi, H; Iwasawa, Y; Okamoto, O; Okuda, S; Ohta, H; Sagara, T Identification of a novel spiropiperidine opioid receptor-like 1 antagonist class by a focused library approach featuring 3D-pharmacophore similarity. J Med Chem49:847-9 (2006) [PubMed] Article Goto, Y; Arai-Otsuki, S; Tachibana, Y; Ichikawa, D; Ozaki, S; Takahashi, H; Iwasawa, Y; Okamoto, O; Okuda, S; Ohta, H; Sagara, T Identification of a novel spiropiperidine opioid receptor-like 1 antagonist class by a focused library approach featuring 3D-pharmacophore similarity. J Med Chem49:847-9 (2006) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Nociceptin receptor |
|---|
| Name: | Nociceptin receptor |
|---|
| Synonyms: | KOR-3 | Kappa-type 3 opioid receptor | Mu-type opioid receptor (Mu) | NOP | Nociceptin Receptor (ORL1 Receptor) | Nociceptin receptor (NOP) | Nociceptin receptor (ORL-1) | Nociceptin receptor (ORL1) | Nociceptin/Orphanin FQ, NOP receptor | OOR | OPIATE ORL-1 | OPRL1 | OPRL1 protein | OPRX_HUMAN | ORL1 | ORL1 receptor | Opioid receptor like-1 | Orphanin FQ receptor | Orphanin FQ receptor (ORL1) | P41146 |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 40702.87 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P41146 |
|---|
| Residue: | 370 |
|---|
| Sequence: | MEPLFPAPFWEVIYGSHLQGNLSLLSPNHSLLPPHLLLNASHGAFLPLGLKVTIVGLYLA
VCVGGLLGNCLVMYVILRHTKMKTATNIYIFNLALADTLVLLTLPFQGTDILLGFWPFGN
ALCKTVIAIDYYNMFTSTFTLTAMSVDRYVAICHPIRALDVRTSSKAQAVNVAIWALASV
VGVPVAIMGSAQVEDEEIECLVEIPTPQDYWGPVFAICIFLFSFIVPVLVISVCYSLMIR
RLRGVRLLSGSREKDRNLRRITRLVLVVVAVFVGCWTPVQVFVLAQGLGVQPSSETAVAI
LRFCTALGYVNSCLNPILYAFLDENFKACFRKFCCASALRRDVQVSDRVRSIAKDVALAC
KTSETVPRPA
|
|
|
|---|
| BDBM50181392 |
|---|
| n/a |
|---|
| Name | BDBM50181392 |
|---|
| Synonyms: | (R)-N-(3-(3H-spiro[isobenzofuran-1,4'-piperidine]-1'-yl)propyl)-1-benzylpyrrolidine-2-carboxamide | 1-benzyl-N-[3-spiro[isobenzofuran-1(3H),4'-piperidine]-1-yl]propyl-D-prolineamide | CHEMBL201945 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C27H35N3O2 |
|---|
| Mol. Mass. | 433.5857 |
|---|
| SMILES | O=C(NCCCN1CCC2(CC1)OCc1ccccc21)[C@H]1CCCN1Cc1ccccc1 |
|---|
| Structure | 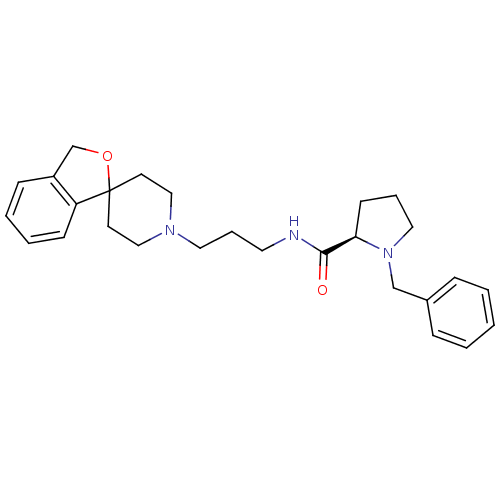
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Goto, Y; Arai-Otsuki, S; Tachibana, Y; Ichikawa, D; Ozaki, S; Takahashi, H; Iwasawa, Y; Okamoto, O; Okuda, S; Ohta, H; Sagara, T Identification of a novel spiropiperidine opioid receptor-like 1 antagonist class by a focused library approach featuring 3D-pharmacophore similarity. J Med Chem49:847-9 (2006) [PubMed] Article
Goto, Y; Arai-Otsuki, S; Tachibana, Y; Ichikawa, D; Ozaki, S; Takahashi, H; Iwasawa, Y; Okamoto, O; Okuda, S; Ohta, H; Sagara, T Identification of a novel spiropiperidine opioid receptor-like 1 antagonist class by a focused library approach featuring 3D-pharmacophore similarity. J Med Chem49:847-9 (2006) [PubMed] Article