| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cannabinoid receptor 1 |
|---|
| Ligand | BDBM50213596 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_459063 (CHEMBL925155) |
|---|
| Ki | 666.4±n/a nM |
|---|
| Citation |  Durdagi, S; Kapou, A; Kourouli, T; Andreou, T; Nikas, SP; Nahmias, VR; Papahatjis, DP; Papadopoulos, MG; Mavromoustakos, T The application of 3D-QSAR studies for novel cannabinoid ligands substituted at the C1' position of the alkyl side chain on the structural requirements for binding to cannabinoid receptors CB1 and CB2. J Med Chem50:2875-85 (2007) [PubMed] Article Durdagi, S; Kapou, A; Kourouli, T; Andreou, T; Nikas, SP; Nahmias, VR; Papahatjis, DP; Papadopoulos, MG; Mavromoustakos, T The application of 3D-QSAR studies for novel cannabinoid ligands substituted at the C1' position of the alkyl side chain on the structural requirements for binding to cannabinoid receptors CB1 and CB2. J Med Chem50:2875-85 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Cannabinoid receptor 1 |
|---|
| Name: | Cannabinoid receptor 1 |
|---|
| Synonyms: | CANN6 | CANNABINOID CB1 | CB-R | CB1 | CNR | CNR1 | CNR1_HUMAN | Cannabinoid CB1 receptor | Cannabinoid receptor | Cannabinoid receptor 1 (CB1) | Cannabinoid receptor 1 (brain) |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 52868.96 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P21554 |
|---|
| Residue: | 472 |
|---|
| Sequence: | MKSILDGLADTTFRTITTDLLYVGSNDIQYEDIKGDMASKLGYFPQKFPLTSFRGSPFQE
KMTAGDNPQLVPADQVNITEFYNKSLSSFKENEENIQCGENFMDIECFMVLNPSQQLAIA
VLSLTLGTFTVLENLLVLCVILHSRSLRCRPSYHFIGSLAVADLLGSVIFVYSFIDFHVF
HRKDSRNVFLFKLGGVTASFTASVGSLFLTAIDRYISIHRPLAYKRIVTRPKAVVAFCLM
WTIAIVIAVLPLLGWNCEKLQSVCSDIFPHIDETYLMFWIGVTSVLLLFIVYAYMYILWK
AHSHAVRMIQRGTQKSIIIHTSEDGKVQVTRPDQARMDIRLAKTLVLILVVLIICWGPLL
AIMVYDVFGKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMFPSCEGTAQ
PLDNSMGDSDCLHKHANNAASVHRAAESCIKSTVKIAKVTMSVSTDTSAEAL
|
|
|
|---|
| BDBM50213596 |
|---|
| n/a |
|---|
| Name | BDBM50213596 |
|---|
| Synonyms: | 5-(2,2-dichloro-1-hexyl-cyclopropyl)-2-((R)-6-isopropenyl-3-methyl-cyclohex-2-enyl)-benzene-1,3-diol | 5-(2,2-dichloro-1-hexylcyclopropyl)-2-((1R)-3-methyl-6-(prop-1-en-2-yl)cyclohex-2-enyl)benzene-1,3-diol | CHEMBL238972 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C25H34Cl2O2 |
|---|
| Mol. Mass. | 437.442 |
|---|
| SMILES | CCCCCCC1(CC1(Cl)Cl)c1cc(O)c([C@@H]2C=C(C)CCC2C(C)=C)c(O)c1 |w:22.24,6.5,t:18| |
|---|
| Structure | 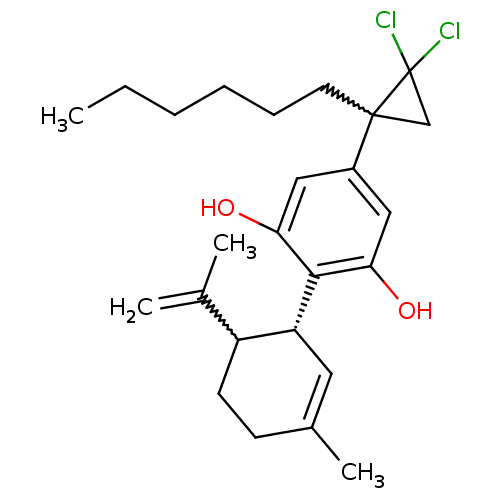
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Durdagi, S; Kapou, A; Kourouli, T; Andreou, T; Nikas, SP; Nahmias, VR; Papahatjis, DP; Papadopoulos, MG; Mavromoustakos, T The application of 3D-QSAR studies for novel cannabinoid ligands substituted at the C1' position of the alkyl side chain on the structural requirements for binding to cannabinoid receptors CB1 and CB2. J Med Chem50:2875-85 (2007) [PubMed] Article
Durdagi, S; Kapou, A; Kourouli, T; Andreou, T; Nikas, SP; Nahmias, VR; Papahatjis, DP; Papadopoulos, MG; Mavromoustakos, T The application of 3D-QSAR studies for novel cannabinoid ligands substituted at the C1' position of the alkyl side chain on the structural requirements for binding to cannabinoid receptors CB1 and CB2. J Med Chem50:2875-85 (2007) [PubMed] Article