| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Serine/threonine-protein kinase MARK2 |
|---|
| Ligand | BDBM50218709 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_444998 (CHEMBL894156) |
|---|
| IC50 | >10000±n/a nM |
|---|
| Citation |  Wang, L; Sullivan, GM; Hexamer, LA; Hasvold, LA; Thalji, R; Przytulinska, M; Tao, ZF; Li, G; Chen, Z; Xiao, Z; Gu, WZ; Xue, J; Bui, MH; Merta, P; Kovar, P; Bouska, JJ; Zhang, H; Park, C; Stewart, KD; Sham, HL; Sowin, TJ; Rosenberg, SH; Lin, NH Design, synthesis, and biological activity of 5,10-dihydro-dibenzo[b,e][1,4]diazepin-11-one-based potent and selective Chk-1 inhibitors. J Med Chem50:4162-76 (2007) [PubMed] Article Wang, L; Sullivan, GM; Hexamer, LA; Hasvold, LA; Thalji, R; Przytulinska, M; Tao, ZF; Li, G; Chen, Z; Xiao, Z; Gu, WZ; Xue, J; Bui, MH; Merta, P; Kovar, P; Bouska, JJ; Zhang, H; Park, C; Stewart, KD; Sham, HL; Sowin, TJ; Rosenberg, SH; Lin, NH Design, synthesis, and biological activity of 5,10-dihydro-dibenzo[b,e][1,4]diazepin-11-one-based potent and selective Chk-1 inhibitors. J Med Chem50:4162-76 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Serine/threonine-protein kinase MARK2 |
|---|
| Name: | Serine/threonine-protein kinase MARK2 |
|---|
| Synonyms: | ELKL motif kinase 1 | EMK1 | MAP/microtubule affinity-regulating kinase 2 | MARK2 | MARK2_HUMAN | PAR1 homolog | Par-1 | Serine/threonine-protein kinase MARK2 |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 87953.09 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ChEMBL_774586 |
|---|
| Residue: | 788 |
|---|
| Sequence: | MSSARTPLPTLNERDTEQPTLGHLDSKPSSKSNMIRGRNSATSADEQPHIGNYRLLKTIG
KGNFAKVKLARHILTGKEVAVKIIDKTQLNSSSLQKLFREVRIMKVLNHPNIVKLFEVIE
TEKTLYLVMEYASGGEVFDYLVAHGRMKEKEARAKFRQIVSAVQYCHQKFIVHRDLKAEN
LLLDADMNIKIADFGFSNEFTFGNKLDTFCGSPPYAAPELFQGKKYDGPEVDVWSLGVIL
YTLVSGSLPFDGQNLKELRERVLRGKYRIPFYMSTDCENLLKKFLILNPSKRGTLEQIMK
DRWMNVGHEDDELKPYVEPLPDYKDPRRTELMVSMGYTREEIQDSLVGQRYNEVMATYLL
LGYKSSELEGDTITLKPRPSADLTNSSAPSPSHKVQRSVSANPKQRRFSDQAAGPAIPTS
NSYSKKTQSNNAENKRPEEDRESGRKASSTAKVPASPLPGLERKKTTPTPSTNSVLSTST
NRSRNSPLLERASLGQASIQNGKDSLTMPGSRASTASASAAVSAARPRQHQKSMSASVHP
NKASGLPPTESNCEVPRPSTAPQRVPVASPSAHNISSSGGAPDRTNFPRGVSSRSTFHAG
QLRQVRDQQNLPYGVTPASPSGHSQGRRGASGSIFSKFTSKFVRRNLSFRFARRNLNEPE
SKDRVETLRPHVVGSGGNDKEKEEFREAKPRSLRFTWSMKTTSSMEPNEMMREIRKVLDA
NSCQSELHEKYMLLCMHGTPGHEDFVQWEMEVCKLPRLSLNGVRFKRISGTSMAFKNIAS
KIANELKL
|
|
|
|---|
| BDBM50218709 |
|---|
| n/a |
|---|
| Name | BDBM50218709 |
|---|
| Synonyms: | 2-[3-(3-methoxy-4-nitrophenyl)-11-oxo-10,11-dihydro-5Hdibenzo[b,e][1,4]diazepin-8-yl]-2-methyl-N-(4-morpholin-4-ylphenyl)propanamide | CHEMBL389745 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C34H35N5O4 |
|---|
| Mol. Mass. | 577.6728 |
|---|
| SMILES | COc1cc(ccc1N)-c1ccc2c(Nc3ccc(cc3NC2=O)C(C)(C)C(=O)Nc2ccc(cc2)N2CCOCC2)c1 |
|---|
| Structure | 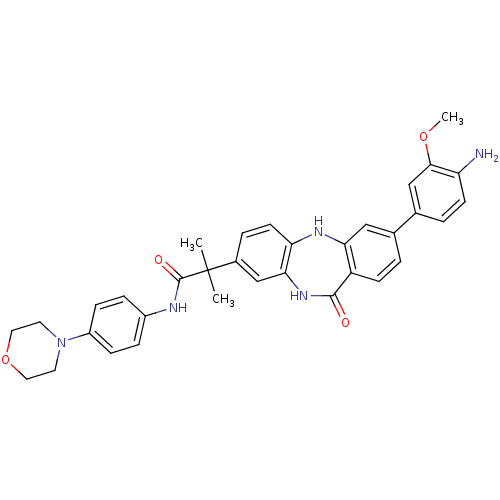
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Wang, L; Sullivan, GM; Hexamer, LA; Hasvold, LA; Thalji, R; Przytulinska, M; Tao, ZF; Li, G; Chen, Z; Xiao, Z; Gu, WZ; Xue, J; Bui, MH; Merta, P; Kovar, P; Bouska, JJ; Zhang, H; Park, C; Stewart, KD; Sham, HL; Sowin, TJ; Rosenberg, SH; Lin, NH Design, synthesis, and biological activity of 5,10-dihydro-dibenzo[b,e][1,4]diazepin-11-one-based potent and selective Chk-1 inhibitors. J Med Chem50:4162-76 (2007) [PubMed] Article
Wang, L; Sullivan, GM; Hexamer, LA; Hasvold, LA; Thalji, R; Przytulinska, M; Tao, ZF; Li, G; Chen, Z; Xiao, Z; Gu, WZ; Xue, J; Bui, MH; Merta, P; Kovar, P; Bouska, JJ; Zhang, H; Park, C; Stewart, KD; Sham, HL; Sowin, TJ; Rosenberg, SH; Lin, NH Design, synthesis, and biological activity of 5,10-dihydro-dibenzo[b,e][1,4]diazepin-11-one-based potent and selective Chk-1 inhibitors. J Med Chem50:4162-76 (2007) [PubMed] Article