| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cannabinoid receptor 2 |
|---|
| Ligand | BDBM50226179 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_451877 (CHEMBL901032) |
|---|
| Ki | 24500±n/a nM |
|---|
| Citation |  Srivastava, BK; Joharapurkar, A; Raval, S; Patel, JZ; Soni, R; Raval, P; Gite, A; Goswami, A; Sadhwani, N; Gandhi, N; Patel, H; Mishra, B; Solanki, M; Pandey, B; Jain, MR; Patel, PR Diaryl dihydropyrazole-3-carboxamides with significant in vivo antiobesity activity related to CB1 receptor antagonism: synthesis, biological evaluation, and molecular modeling in the homology model. J Med Chem50:5951-66 (2007) [PubMed] Article Srivastava, BK; Joharapurkar, A; Raval, S; Patel, JZ; Soni, R; Raval, P; Gite, A; Goswami, A; Sadhwani, N; Gandhi, N; Patel, H; Mishra, B; Solanki, M; Pandey, B; Jain, MR; Patel, PR Diaryl dihydropyrazole-3-carboxamides with significant in vivo antiobesity activity related to CB1 receptor antagonism: synthesis, biological evaluation, and molecular modeling in the homology model. J Med Chem50:5951-66 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Cannabinoid receptor 2 |
|---|
| Name: | Cannabinoid receptor 2 |
|---|
| Synonyms: | CANNABINOID CB2 | CB-2 | CB2 | CB2A | CB2B | CNR2 | CNR2_HUMAN | CX5 | Cannabinoid CB2 receptor | Cannabinoid receptor 2 (CB2) | Cannabinoid receptor 2 (CB2R) | hCB2 |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 39690.94 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P34972 |
|---|
| Residue: | 360 |
|---|
| Sequence: | MEECWVTEIANGSKDGLDSNPMKDYMILSGPQKTAVAVLCTLLGLLSALENVAVLYLILS
SHQLRRKPSYLFIGSLAGADFLASVVFACSFVNFHVFHGVDSKAVFLLKIGSVTMTFTAS
VGSLLLTAIDRYLCLRYPPSYKALLTRGRALVTLGIMWVLSALVSYLPLMGWTCCPRPCS
ELFPLIPNDYLLSWLLFIAFLFSGIIYTYGHVLWKAHQHVASLSGHQDRQVPGMARMRLD
VRLAKTLGLVLAVLLICWFPVLALMAHSLATTLSDQVKKAFAFCSMLCLINSMVNPVIYA
LRSGEIRSSAHHCLAHWKKCVRGLGSEAKEEAPRSSVTETEADGKITPWPDSRDLDLSDC
|
|
|
|---|
| BDBM50226179 |
|---|
| n/a |
|---|
| Name | BDBM50226179 |
|---|
| Synonyms: | (+)-5-(4-chlorophenyl)-1-(2,4-dichlorophenyl)-4,5-dihydro-1H-pyrazole-3-carboxylic acid morpholin-4-ylamide bisulphate salt | (-)-5-(4-chlorophenyl)-1-(2,4-dichlorophenyl)-4,5-dihydro-1H-pyrazole-3-carboxylic acid morpholin-4-ylamide bisulphate salt | 5-(4-chlorophenyl)-1-(2,4-dichlorophenyl)-4,5-dihydro-1H-pyrazole-3-carboxylic acid morpholin-4-ylamide bisulphate salt | CHEMBL239703 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H19Cl3N4O2 |
|---|
| Mol. Mass. | 453.749 |
|---|
| SMILES | Clc1ccc(cc1)C1CC(=NN1c1ccc(Cl)cc1Cl)C(=O)NN1CCOCC1 |c:10| |
|---|
| Structure | 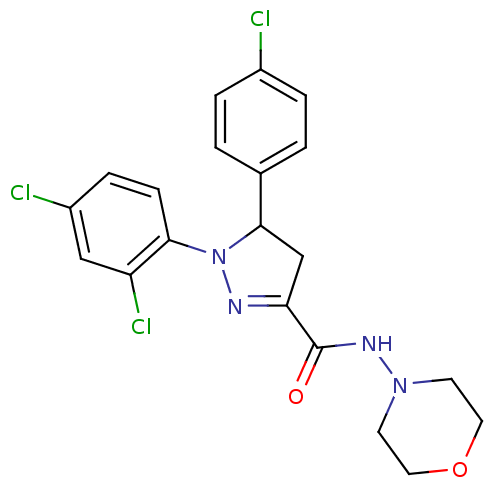
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Srivastava, BK; Joharapurkar, A; Raval, S; Patel, JZ; Soni, R; Raval, P; Gite, A; Goswami, A; Sadhwani, N; Gandhi, N; Patel, H; Mishra, B; Solanki, M; Pandey, B; Jain, MR; Patel, PR Diaryl dihydropyrazole-3-carboxamides with significant in vivo antiobesity activity related to CB1 receptor antagonism: synthesis, biological evaluation, and molecular modeling in the homology model. J Med Chem50:5951-66 (2007) [PubMed] Article
Srivastava, BK; Joharapurkar, A; Raval, S; Patel, JZ; Soni, R; Raval, P; Gite, A; Goswami, A; Sadhwani, N; Gandhi, N; Patel, H; Mishra, B; Solanki, M; Pandey, B; Jain, MR; Patel, PR Diaryl dihydropyrazole-3-carboxamides with significant in vivo antiobesity activity related to CB1 receptor antagonism: synthesis, biological evaluation, and molecular modeling in the homology model. J Med Chem50:5951-66 (2007) [PubMed] Article