| Citation |  Glossop, PA; Watson, CA; Price, DA; Bunnage, ME; Middleton, DS; Wood, A; James, K; Roberts, D; Strang, RS; Yeadon, M; Perros-Huguet, C; Clarke, NP; Trevethick, MA; Machin, I; Stuart, EF; Evans, SM; Harrison, AC; Fairman, DA; Agoram, B; Burrows, JL; Feeder, N; Fulton, CK; Dillon, BR; Entwistle, DA; Spence, FJ Inhalation by design: novel tertiary amine muscarinic M3 receptor antagonists with slow off-rate binding kinetics for inhaled once-daily treatment of chronic obstructive pulmonary disease. J Med Chem54:6888-904 (2011) [PubMed] Article Glossop, PA; Watson, CA; Price, DA; Bunnage, ME; Middleton, DS; Wood, A; James, K; Roberts, D; Strang, RS; Yeadon, M; Perros-Huguet, C; Clarke, NP; Trevethick, MA; Machin, I; Stuart, EF; Evans, SM; Harrison, AC; Fairman, DA; Agoram, B; Burrows, JL; Feeder, N; Fulton, CK; Dillon, BR; Entwistle, DA; Spence, FJ Inhalation by design: novel tertiary amine muscarinic M3 receptor antagonists with slow off-rate binding kinetics for inhaled once-daily treatment of chronic obstructive pulmonary disease. J Med Chem54:6888-904 (2011) [PubMed] Article |
|---|
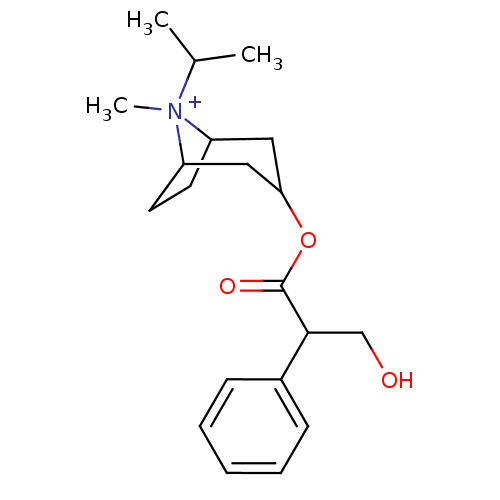
| SMILES | CC(C)[N+]1(C)C2CCC1CC(C2)OC(=O)C(CO)c1ccccc1 |TLB:4:3:6.7:11.9.10,1:3:6.7:11.9.10,(10.09,.76,;9.35,2.05,;7.85,2.05,;10.09,3.35,;11.39,2.6,;10.09,4.89,;10.86,3.55,;10.3,2.58,;8.76,2.58,;7.43,3.35,;7.43,4.89,;8.76,5.66,;6.09,5.66,;6.09,7.2,;7.43,7.97,;4.76,7.97,;4.76,9.51,;3.43,10.28,;3.43,7.2,;3.43,5.66,;2.09,4.89,;.76,5.66,;.76,7.2,;2.09,7.97,)| |
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Glossop, PA; Watson, CA; Price, DA; Bunnage, ME; Middleton, DS; Wood, A; James, K; Roberts, D; Strang, RS; Yeadon, M; Perros-Huguet, C; Clarke, NP; Trevethick, MA; Machin, I; Stuart, EF; Evans, SM; Harrison, AC; Fairman, DA; Agoram, B; Burrows, JL; Feeder, N; Fulton, CK; Dillon, BR; Entwistle, DA; Spence, FJ Inhalation by design: novel tertiary amine muscarinic M3 receptor antagonists with slow off-rate binding kinetics for inhaled once-daily treatment of chronic obstructive pulmonary disease. J Med Chem54:6888-904 (2011) [PubMed] Article
Glossop, PA; Watson, CA; Price, DA; Bunnage, ME; Middleton, DS; Wood, A; James, K; Roberts, D; Strang, RS; Yeadon, M; Perros-Huguet, C; Clarke, NP; Trevethick, MA; Machin, I; Stuart, EF; Evans, SM; Harrison, AC; Fairman, DA; Agoram, B; Burrows, JL; Feeder, N; Fulton, CK; Dillon, BR; Entwistle, DA; Spence, FJ Inhalation by design: novel tertiary amine muscarinic M3 receptor antagonists with slow off-rate binding kinetics for inhaled once-daily treatment of chronic obstructive pulmonary disease. J Med Chem54:6888-904 (2011) [PubMed] Article