

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Muscarinic acetylcholine receptor M1 | ||
| Ligand | BDBM50033162 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_138811 (CHEMBL747492) | ||
| IC50 | 0.70±n/a nM | ||
| Citation |  Ward, JS; Merritt, L; Calligaro, DO; Bymaster, FP; Shannon, HE; Sawyer, BD; Mitch, CH; Deeter, JB; Peters, SC; Sheardown, MJ Functionally selective M1 muscarinic agonists. 3. Side chains and azacycles contributing to functional muscarinic selectivity among pyrazinylazacycles. J Med Chem38:3469-81 (1995) [PubMed] Ward, JS; Merritt, L; Calligaro, DO; Bymaster, FP; Shannon, HE; Sawyer, BD; Mitch, CH; Deeter, JB; Peters, SC; Sheardown, MJ Functionally selective M1 muscarinic agonists. 3. Side chains and azacycles contributing to functional muscarinic selectivity among pyrazinylazacycles. J Med Chem38:3469-81 (1995) [PubMed] | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Muscarinic acetylcholine receptor M1 | |||
| Name: | Muscarinic acetylcholine receptor M1 | ||
| Synonyms: | ACM1_RAT | Cholinergic, muscarinic M1 | Chrm-1 | Chrm1 | cholinergic receptor, muscarinic 1 | ||
| Type: | Enzyme Catalytic Domain | ||
| Mol. Mass.: | 51390.46 | ||
| Organism: | RAT | ||
| Description: | P08482 | ||
| Residue: | 460 | ||
| Sequence: |
| ||
| BDBM50033162 | |||
| n/a | |||
| Name | BDBM50033162 | ||
| Synonyms: | 3-(3-Pentylsulfanyl-pyrazin-2-yl)-1-aza-bicyclo[2.2.2]octane | CHEMBL331805 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C16H25N3S | ||
| Mol. Mass. | 291.455 | ||
| SMILES | CCCCCSc1nccnc1C1CN2CCC1CC2 |(5.44,2.49,;5.44,.95,;6.77,.18,;6.77,-1.36,;8.1,-2.13,;8.1,-3.67,;9.43,-4.44,;10.76,-3.69,;12.09,-4.46,;12.09,-6,;10.76,-6.75,;9.43,-5.98,;8.1,-6.75,;8.1,-8.29,;6.77,-9.05,;7.31,-7.51,;6.03,-7.4,;6.77,-5.97,;5.44,-6.75,;5.44,-8.29,)| | ||
| Structure | 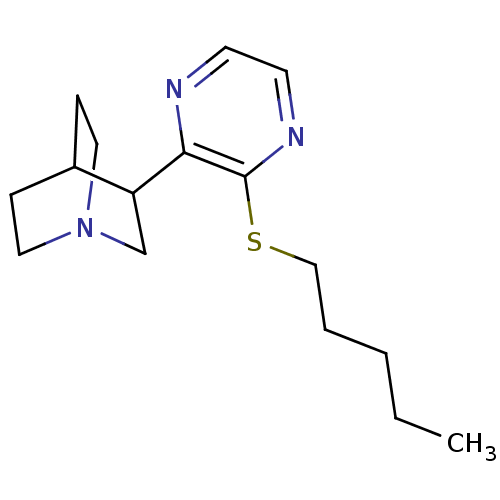
| ||