| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Mitogen-activated protein kinase 10 |
|---|
| Ligand | BDBM50303636 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_596583 (CHEMBL1042602) |
|---|
| IC50 | 491±n/a nM |
|---|
| Citation |  Kamenecka, T; Jiang, R; Song, X; Duckett, D; Chen, W; Ling, YY; Habel, J; Laughlin, JD; Chambers, J; Figuera-Losada, M; Cameron, MD; Lin, L; Ruiz, CH; LoGrasso, PV Synthesis, biological evaluation, X-ray structure, and pharmacokinetics of aminopyrimidine c-jun-N-terminal kinase (JNK) inhibitors. J Med Chem53:419-31 (2010) [PubMed] Article Kamenecka, T; Jiang, R; Song, X; Duckett, D; Chen, W; Ling, YY; Habel, J; Laughlin, JD; Chambers, J; Figuera-Losada, M; Cameron, MD; Lin, L; Ruiz, CH; LoGrasso, PV Synthesis, biological evaluation, X-ray structure, and pharmacokinetics of aminopyrimidine c-jun-N-terminal kinase (JNK) inhibitors. J Med Chem53:419-31 (2010) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Mitogen-activated protein kinase 10 |
|---|
| Name: | Mitogen-activated protein kinase 10 |
|---|
| Synonyms: | JNK3 | JNK3A | MAP kinase p49 3F12 | MAPK10 | MK10_HUMAN | Mitogen-Activated Protein Kinase 10 (JNK3) | Mitogen-activated protein kinase 10 (Stress-activated protein kinase JNK3) (c-Jun N-terminal kinase 3) (MAP kinase p49 3F12) | Mitogen-activated protein kinase 10/Receptor-interacting serine/threonine-protein kinase 1 | PRKM10 | SAPK1B | Stress-activated protein kinase JNK3 | c-Jun N-terminal kinase 3 (JNK3) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 52586.89 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 464 |
|---|
| Sequence: | MSLHFLYYCSEPTLDVKIAFCQGFDKQVDVSYIAKHYNMSKSKVDNQFYSVEVGDSTFTV
LKRYQNLKPIGSGAQGIVCAAYDAVLDRNVAIKKLSRPFQNQTHAKRAYRELVLMKCVNH
KNIISLLNVFTPQKTLEEFQDVYLVMELMDANLCQVIQMELDHERMSYLLYQMLCGIKHL
HSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTAGTSFMMTPYVVTRYYRAPEVILGMGY
KENVDIWSVGCIMGEMVRHKILFPGRDYIDQWNKVIEQLGTPCPEFMKKLQPTVRNYVEN
RPKYAGLTFPKLFPDSLFPADSEHNKLKASQARDLLSKMLVIDPAKRISVDDALQHPYIN
VWYDPAEVEAPPPQIYDKQLDEREHTIEEWKELIYKEVMNSEEKTKNGVVKGQPSPSGAA
VNSSESLPPSSSVNDISSMSTDQTLASDTDSSLEASAGPLGCCR
|
|
|
|---|
| BDBM50303636 |
|---|
| n/a |
|---|
| Name | BDBM50303636 |
|---|
| Synonyms: | 4-(4-Phenylpyrimidin-2-ylamino)benzamide | CHEMBL571619 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C17H14N4O |
|---|
| Mol. Mass. | 290.3193 |
|---|
| SMILES | NC(=O)c1ccc(Nc2nccc(n2)-c2ccccc2)cc1 |
|---|
| Structure | 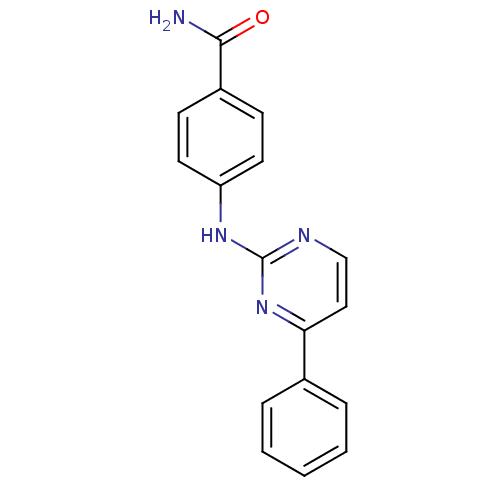
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Kamenecka, T; Jiang, R; Song, X; Duckett, D; Chen, W; Ling, YY; Habel, J; Laughlin, JD; Chambers, J; Figuera-Losada, M; Cameron, MD; Lin, L; Ruiz, CH; LoGrasso, PV Synthesis, biological evaluation, X-ray structure, and pharmacokinetics of aminopyrimidine c-jun-N-terminal kinase (JNK) inhibitors. J Med Chem53:419-31 (2010) [PubMed] Article
Kamenecka, T; Jiang, R; Song, X; Duckett, D; Chen, W; Ling, YY; Habel, J; Laughlin, JD; Chambers, J; Figuera-Losada, M; Cameron, MD; Lin, L; Ruiz, CH; LoGrasso, PV Synthesis, biological evaluation, X-ray structure, and pharmacokinetics of aminopyrimidine c-jun-N-terminal kinase (JNK) inhibitors. J Med Chem53:419-31 (2010) [PubMed] Article