

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Cytochrome P450 2C9 | ||
| Ligand | BDBM50380677 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_813215 (CHEMBL2020947) | ||
| IC50 | 2300±n/a nM | ||
| Citation |  Mirguet, O; Lamotte, Y; Donche, F; Toum, J; Gellibert, F; Bouillot, A; Gosmini, R; Nguyen, VL; Delannée, D; Seal, J; Blandel, F; Boullay, AB; Boursier, E; Martin, S; Brusq, JM; Krysa, G; Riou, A; Tellier, R; Costaz, A; Huet, P; Dudit, Y; Trottet, L; Kirilovsky, J; Nicodeme, E From ApoA1 upregulation to BET family bromodomain inhibition: discovery of I-BET151. Bioorg Med Chem Lett22:2963-7 (2012) [PubMed] Article Mirguet, O; Lamotte, Y; Donche, F; Toum, J; Gellibert, F; Bouillot, A; Gosmini, R; Nguyen, VL; Delannée, D; Seal, J; Blandel, F; Boullay, AB; Boursier, E; Martin, S; Brusq, JM; Krysa, G; Riou, A; Tellier, R; Costaz, A; Huet, P; Dudit, Y; Trottet, L; Kirilovsky, J; Nicodeme, E From ApoA1 upregulation to BET family bromodomain inhibition: discovery of I-BET151. Bioorg Med Chem Lett22:2963-7 (2012) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Cytochrome P450 2C9 | |||
| Name: | Cytochrome P450 2C9 | ||
| Synonyms: | (R)-limonene 6-monooxygenase | (S)-limonene 6-monooxygenase | CP2C9_HUMAN | CYP2C10 | CYP2C9 | CYPIIC9 | Cytochrome P450 2C9 (CYP2C9 ) | Cytochrome P450 2C9 (CYP2C9) | P-450MP | P450 MP-4/MP-8 | P450 PB-1 | S-mephenytoin 4-hydroxylase | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 55636.33 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P11712 | ||
| Residue: | 490 | ||
| Sequence: |
| ||
| BDBM50380677 | |||
| n/a | |||
| Name | BDBM50380677 | ||
| Synonyms: | CHEMBL2017284 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C23H17F3N4O4 | ||
| Mol. Mass. | 470.4007 | ||
| SMILES | COc1cc2c3n(-c4ccccc4OC(F)(F)F)c(=O)[nH]c3cnc2cc1-c1c(C)noc1C |(17.24,-27.14,;17.24,-28.68,;18.57,-29.45,;19.9,-28.68,;21.23,-29.44,;22.56,-28.67,;22.87,-27.17,;22.1,-25.83,;20.56,-25.84,;19.79,-24.51,;20.56,-23.17,;22.1,-23.17,;22.87,-24.5,;24.41,-24.51,;25.18,-23.18,;26.72,-23.18,;24.42,-21.84,;25.94,-21.83,;24.39,-26.99,;25.15,-25.65,;25.03,-28.39,;23.9,-29.42,;23.91,-30.98,;22.57,-31.75,;21.24,-30.98,;19.9,-31.76,;18.57,-30.99,;17.23,-31.76,;17.2,-33.3,;18.43,-34.23,;15.73,-33.75,;14.85,-32.48,;15.78,-31.26,;15.33,-29.78,)| | ||
| Structure | 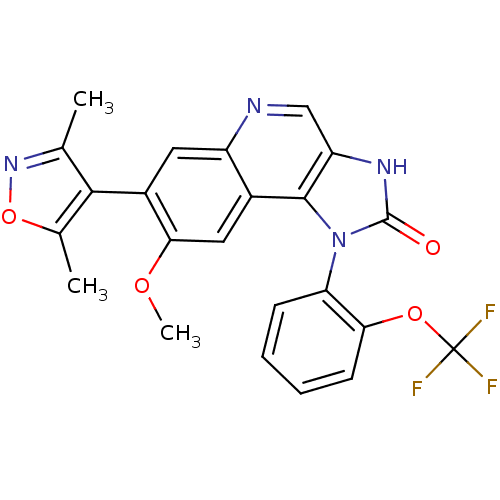
| ||