| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform |
|---|
| Ligand | BDBM50394174 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_856505 (CHEMBL2161551) |
|---|
| IC50 | 93±n/a nM |
|---|
| Citation |  Gopalsamy, A; Bennett, EM; Shi, M; Zhang, WG; Bard, J; Yu, K Identification of pyrimidine derivatives as hSMG-1 inhibitors. Bioorg Med Chem Lett22:6636-41 (2012) [PubMed] Article Gopalsamy, A; Bennett, EM; Shi, M; Zhang, WG; Bard, J; Yu, K Identification of pyrimidine derivatives as hSMG-1 inhibitors. Bioorg Med Chem Lett22:6636-41 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform |
|---|
| Name: | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform |
|---|
| Synonyms: | PI3-kinase p110 subunit gamma | PI3-kinase subunit p120-gamma | PI3Kgamma | PIK3CG | PK3CG_HUMAN | Phosphatidylinositol 4,5-biphosphate 3-kinase catalytic subunit gamma (PIK3CG) | Phosphatidylinositol 4,5-bisphosphate 3-kinase (PI3K) | Phosphatidylinositol 4,5-bisphosphate 3-kinase 110 kDa catalytic subunit gamma (PI3K gamma) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma (PI3Kgamma) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (PI3K gamma) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (PI3K) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (PI3Kgamma) | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | Phosphoinositide 3-Kinase (PI3K), gamma Chain A | Phosphoinositide 3-kinases gamma (PI3K gamma) | Phosphoinositide-3-kinase (PI3K gamma) | p120-PI3K |
|---|
| Type: | Enzyme Subunit |
|---|
| Mol. Mass.: | 126470.30 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P48736 |
|---|
| Residue: | 1102 |
|---|
| Sequence: | MELENYKQPVVLREDNCRRRRRMKPRSAAASLSSMELIPIEFVLPTSQRKCKSPETALLH
VAGHGNVEQMKAQVWLRALETSVAADFYHRLGPHHFLLLYQKKGQWYEIYDKYQVVQTLD
CLRYWKATHRSPGQIHLVQRHPPSEESQAFQRQLTALIGYDVTDVSNVHDDELEFTRRGL
VTPRMAEVASRDPKLYAMHPWVTSKPLPEYLWKKIANNCIFIVIHRSTTSQTIKVSPDDT
PGAILQSFFTKMAKKKSLMDIPESQSEQDFVLRVCGRDEYLVGETPIKNFQWVRHCLKNG
EEIHVVLDTPPDPALDEVRKEEWPLVDDCTGVTGYHEQLTIHGKDHESVFTVSLWDCDRK
FRVKIRGIDIPVLPRNTDLTVFVEANIQHGQQVLCQRRTSPKPFTEEVLWNVWLEFSIKI
KDLPKGALLNLQIYCGKAPALSSKASAESPSSESKGKVQLLYYVNLLLIDHRFLLRRGEY
VLHMWQISGKGEDQGSFNADKLTSATNPDKENSMSISILLDNYCHPIALPKHQPTPDPEG
DRVRAEMPNQLRKQLEAIIATDPLNPLTAEDKELLWHFRYESLKHPKAYPKLFSSVKWGQ
QEIVAKTYQLLARREVWDQSALDVGLTMQLLDCNFSDENVRAIAVQKLESLEDDDVLHYL
LQLVQAVKFEPYHDSALARFLLKRGLRNKRIGHFLFWFLRSEIAQSRHYQQRFAVILEAY
LRGCGTAMLHDFTQQVQVIEMLQKVTLDIKSLSAEKYDVSSQVISQLKQKLENLQNSQLP
ESFRVPYDPGLKAGALAIEKCKVMASKKKPLWLEFKCADPTALSNETIGIIFKHGDDLRQ
DMLILQILRIMESIWETESLDLCLLPYGCISTGDKIGMIEIVKDATTIAKIQQSTVGNTG
AFKDEVLNHWLKEKSPTEEKFQAAVERFVYSCAGYCVATFVLGIGDRHNDNIMITETGNL
FHIDFGHILGNYKSFLGINKERVPFVLTPDFLFVMGTSGKKTSPHFQKFQDICVKAYLAL
RHHTNLLIILFSMMLMTGMPQLTSKEDIEYIRDALTVGKNEEDAKKYFLDQIEVCRDKGW
TVQFNWFLHLVLGIKQGEKHSA
|
|
|
|---|
| BDBM50394174 |
|---|
| n/a |
|---|
| Name | BDBM50394174 |
|---|
| Synonyms: | CHEMBL2158863 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C28H31N7O3S |
|---|
| Mol. Mass. | 545.656 |
|---|
| SMILES | CCN(CC)S(=O)(=O)c1cc(Nc2nccc(n2)-c2ccnc(c2)-c2ccc(NC(=O)NC)cc2)ccc1C |
|---|
| Structure | 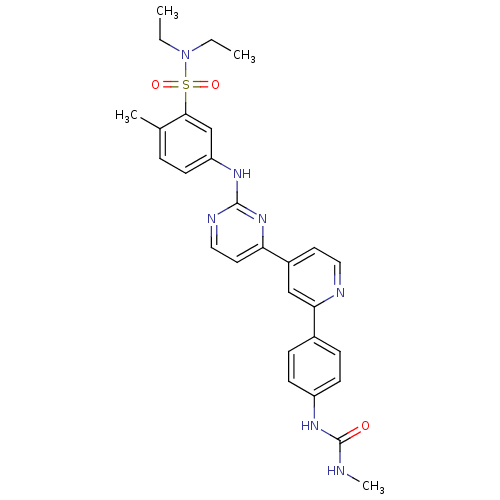
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Gopalsamy, A; Bennett, EM; Shi, M; Zhang, WG; Bard, J; Yu, K Identification of pyrimidine derivatives as hSMG-1 inhibitors. Bioorg Med Chem Lett22:6636-41 (2012) [PubMed] Article
Gopalsamy, A; Bennett, EM; Shi, M; Zhang, WG; Bard, J; Yu, K Identification of pyrimidine derivatives as hSMG-1 inhibitors. Bioorg Med Chem Lett22:6636-41 (2012) [PubMed] Article