| Citation |  Barlind, JG; Bauer, UA; Birch, AM; Birtles, S; Buckett, LK; Butlin, RJ; Davies, RD; Eriksson, JW; Hammond, CD; Hovland, R; Johannesson, P; Johansson, MJ; Kemmitt, PD; Lindmark, BT; Morentin Gutierrez, P; Noeske, TA; Nordin, A; O'Donnell, CJ; Petersson, AU; Redzic, A; Turnbull, AV; Vinblad, J Design and optimization of pyrazinecarboxamide-based inhibitors of diacylglycerol acyltransferase 1 (DGAT1) leading to a clinical candidate dimethylpyrazinecarboxamide phenylcyclohexylacetic acid (AZD7687). J Med Chem55:10610-29 (2012) [PubMed] Article Barlind, JG; Bauer, UA; Birch, AM; Birtles, S; Buckett, LK; Butlin, RJ; Davies, RD; Eriksson, JW; Hammond, CD; Hovland, R; Johannesson, P; Johansson, MJ; Kemmitt, PD; Lindmark, BT; Morentin Gutierrez, P; Noeske, TA; Nordin, A; O'Donnell, CJ; Petersson, AU; Redzic, A; Turnbull, AV; Vinblad, J Design and optimization of pyrazinecarboxamide-based inhibitors of diacylglycerol acyltransferase 1 (DGAT1) leading to a clinical candidate dimethylpyrazinecarboxamide phenylcyclohexylacetic acid (AZD7687). J Med Chem55:10610-29 (2012) [PubMed] Article |
|---|
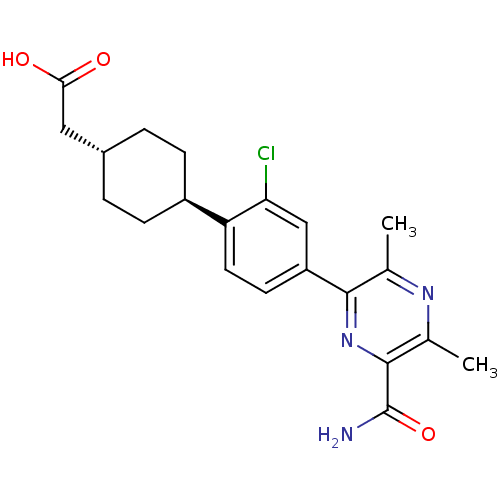
| SMILES | Cc1nc(C)c(nc1C(N)=O)-c1ccc([C@H]2CC[C@H](CC(O)=O)CC2)c(Cl)c1 |r,wU:15.15,wD:18.19,(21.76,-10.54,;23.09,-9.77,;24.44,-10.54,;25.76,-9.77,;27.1,-10.53,;25.76,-8.23,;24.43,-7.46,;23.09,-8.23,;21.76,-7.45,;20.43,-8.22,;21.76,-5.91,;27.09,-7.46,;27.09,-5.92,;28.42,-5.15,;29.76,-5.91,;31.09,-5.14,;31.08,-3.6,;32.42,-2.83,;33.76,-3.6,;35.09,-2.83,;36.42,-3.6,;37.76,-2.83,;36.42,-5.14,;33.75,-5.14,;32.43,-5.91,;29.76,-7.46,;31.1,-8.23,;28.43,-8.23,)| |
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Barlind, JG; Bauer, UA; Birch, AM; Birtles, S; Buckett, LK; Butlin, RJ; Davies, RD; Eriksson, JW; Hammond, CD; Hovland, R; Johannesson, P; Johansson, MJ; Kemmitt, PD; Lindmark, BT; Morentin Gutierrez, P; Noeske, TA; Nordin, A; O'Donnell, CJ; Petersson, AU; Redzic, A; Turnbull, AV; Vinblad, J Design and optimization of pyrazinecarboxamide-based inhibitors of diacylglycerol acyltransferase 1 (DGAT1) leading to a clinical candidate dimethylpyrazinecarboxamide phenylcyclohexylacetic acid (AZD7687). J Med Chem55:10610-29 (2012) [PubMed] Article
Barlind, JG; Bauer, UA; Birch, AM; Birtles, S; Buckett, LK; Butlin, RJ; Davies, RD; Eriksson, JW; Hammond, CD; Hovland, R; Johannesson, P; Johansson, MJ; Kemmitt, PD; Lindmark, BT; Morentin Gutierrez, P; Noeske, TA; Nordin, A; O'Donnell, CJ; Petersson, AU; Redzic, A; Turnbull, AV; Vinblad, J Design and optimization of pyrazinecarboxamide-based inhibitors of diacylglycerol acyltransferase 1 (DGAT1) leading to a clinical candidate dimethylpyrazinecarboxamide phenylcyclohexylacetic acid (AZD7687). J Med Chem55:10610-29 (2012) [PubMed] Article