

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Vitamin K-dependent protein C | ||
| Ligand | BDBM50425661 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_937463 (CHEMBL2320211) | ||
| Ki | 39±n/a nM | ||
| Citation |  Saupe, SM; Leubner, S; Betz, M; Klebe, G; Steinmetzer, T Development of new cyclic plasmin inhibitors with excellent potency and selectivity. J Med Chem56:820-31 (2013) [PubMed] Article Saupe, SM; Leubner, S; Betz, M; Klebe, G; Steinmetzer, T Development of new cyclic plasmin inhibitors with excellent potency and selectivity. J Med Chem56:820-31 (2013) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Vitamin K-dependent protein C | |||
| Name: | Vitamin K-dependent protein C | ||
| Synonyms: | Activated protein C cofactor | Anticoagulant protein C | Apolipoprotein H | Autoprothrombin IIA | Blood coagulation factor XIV | Coagulation factor V | Coagulation factor V heavy chain | Coagulation factor V light chain | Endothelial protein C receptor | PROC | PROC_HUMAN | Proaccelerin, labile factor | Vitamin K-dependent protein C precursor | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 52067.73 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | n/a | ||
| Residue: | 461 | ||
| Sequence: |
| ||
| BDBM50425661 | |||
| n/a | |||
| Name | BDBM50425661 | ||
| Synonyms: | CHEMBL2315240 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C41H47N9O6S | ||
| Mol. Mass. | 793.934 | ||
| SMILES | NC(=N)c1ccc(CNC(=O)[C@@H]2Cc3cccc(NC(=O)CN4CCN(CC4)CC(=O)Nc4cccc(C[C@@H](NS(=O)(=O)Cc5ccccc5)C(=O)N2)c4)c3)cc1 |r,wU:38.39,wD:11.10,(46.63,-3.72,;46.61,-5.26,;47.93,-6.04,;45.27,-6.01,;45.24,-7.55,;43.89,-8.29,;42.58,-7.51,;41.23,-8.26,;39.92,-7.47,;38.57,-8.22,;38.55,-9.76,;37.25,-7.44,;35.9,-8.19,;35.89,-9.73,;34.55,-10.49,;34.54,-12.03,;35.87,-12.8,;37.21,-12.04,;38.54,-12.81,;38.53,-14.34,;39.86,-15.11,;36.03,-17.42,;34.69,-16.66,;33.37,-17.44,;32.04,-16.69,;32.03,-15.16,;33.34,-14.37,;34.68,-15.13,;30.7,-14.41,;30.68,-12.88,;29.34,-12.12,;32,-12.09,;31.98,-10.56,;30.63,-9.8,;30.62,-8.25,;31.95,-7.48,;33.28,-8.24,;34.61,-7.46,;34.59,-5.92,;33.25,-5.16,;31.92,-5.95,;31.15,-7.27,;32.69,-7.27,;30.58,-5.18,;29.26,-5.96,;27.92,-5.19,;26.59,-5.96,;26.59,-7.51,;27.92,-8.28,;29.26,-7.51,;35.92,-5.13,;35.91,-3.59,;37.26,-5.89,;33.3,-9.77,;37.22,-10.5,;42.59,-5.97,;43.93,-5.22,)| | ||
| Structure | 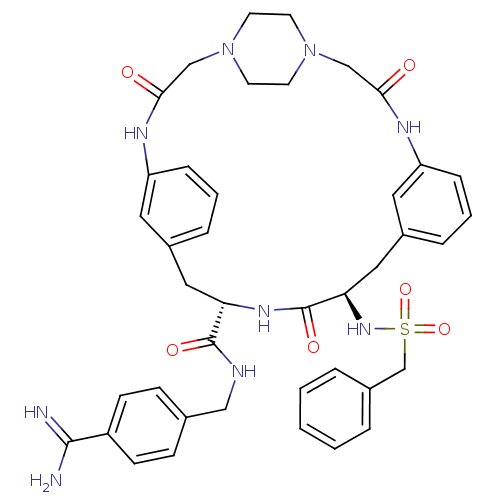
| ||