| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Heat shock protein 75 kDa, mitochondrial |
|---|
| Ligand | BDBM20926 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1284500 (CHEMBL3106548) |
|---|
| Ki | 16±n/a nM |
|---|
| Citation |  Ernst, JT; Liu, M; Zuccola, H; Neubert, T; Beaumont, K; Turnbull, A; Kallel, A; Vought, B; Stamos, D Correlation between chemotype-dependent binding conformations of HSP90a/� and isoform selectivity-Implications for the structure-based design of HSP90a/� selective inhibitors for treating neurodegenerative diseases. Bioorg Med Chem Lett24:204-8 (2013) [PubMed] Article Ernst, JT; Liu, M; Zuccola, H; Neubert, T; Beaumont, K; Turnbull, A; Kallel, A; Vought, B; Stamos, D Correlation between chemotype-dependent binding conformations of HSP90a/� and isoform selectivity-Implications for the structure-based design of HSP90a/� selective inhibitors for treating neurodegenerative diseases. Bioorg Med Chem Lett24:204-8 (2013) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Heat shock protein 75 kDa, mitochondrial |
|---|
| Name: | Heat shock protein 75 kDa, mitochondrial |
|---|
| Synonyms: | HSP75 | TRAP1 | TRAP1_HUMAN |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 80120.13 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ChEMBL_1478164 |
|---|
| Residue: | 704 |
|---|
| Sequence: | MARELRALLLWGRRLRPLLRAPALAAVPGGKPILCPRRTTAQLGPRRNPAWSLQAGRLFS
TQTAEDKEEPLHSIISSTESVQGSTSKHEFQAETKKLLDIVARSLYSEKEVFIRELISNA
SDALEKLRHKLVSDGQALPEMEIHLQTNAEKGTITIQDTGIGMTQEELVSNLGTIARSGS
KAFLDALQNQAEASSKIIGQFGVGFYSAFMVADRVEVYSRSAAPGSLGYQWLSDGSGVFE
IAEASGVRTGTKIIIHLKSDCKEFSSEARVRDVVTKYSNFVSFPLYLNGRRMNTLQAIWM
MDPKDVREWQHEEFYRYVAQAHDKPRYTLHYKTDAPLNIRSIFYVPDMKPSMFDVSRELG
SSVALYSRKVLIQTKATDILPKWLRFIRGVVDSEDIPLNLSRELLQESALIRKLRDVLQQ
RLIKFFIDQSKKDAEKYAKFFEDYGLFMREGIVTATEQEVKEDIAKLLRYESSALPSGQL
TSLSEYASRMRAGTRNIYYLCAPNRHLAEHSPYYEAMKKKDTEVLFCFEQFDELTLLHLR
EFDKKKLISVETDIVVDHYKEEKFEDRSPAAECLSEKETEELMAWMRNVLGSRVTNVKVT
LRLDTHPAMVTVLEMGAARHFLRMQQLAKTQEERAQLLQPTLEINPRHALIKKLNQLRAS
EPGLAQLLVDQIYENAMIAAGLVDDPRAMVGRLNELLVKALERH
|
|
|
|---|
| BDBM20926 |
|---|
| n/a |
|---|
| Name | BDBM20926 |
|---|
| Synonyms: | 5-[2,4-dihydroxy-5-(propan-2-yl)phenyl]-N-ethyl-4-[4-(morpholin-4-ylmethyl)phenyl]-1,2-oxazole-3-carboxamide | Isoxazole, 40f | VER-52296/NVP-AUY922 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H31N3O5 |
|---|
| Mol. Mass. | 465.5414 |
|---|
| SMILES | CCNC(=O)c1noc(c1-c1ccc(CN2CCOCC2)cc1)-c1cc(C(C)C)c(O)cc1O |
|---|
| Structure | 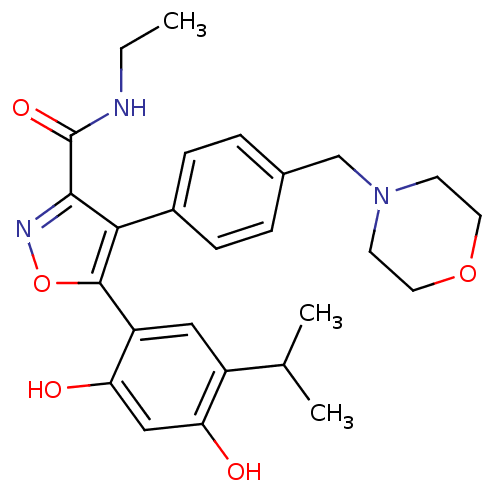
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Ernst, JT; Liu, M; Zuccola, H; Neubert, T; Beaumont, K; Turnbull, A; Kallel, A; Vought, B; Stamos, D Correlation between chemotype-dependent binding conformations of HSP90a/� and isoform selectivity-Implications for the structure-based design of HSP90a/� selective inhibitors for treating neurodegenerative diseases. Bioorg Med Chem Lett24:204-8 (2013) [PubMed] Article
Ernst, JT; Liu, M; Zuccola, H; Neubert, T; Beaumont, K; Turnbull, A; Kallel, A; Vought, B; Stamos, D Correlation between chemotype-dependent binding conformations of HSP90a/� and isoform selectivity-Implications for the structure-based design of HSP90a/� selective inhibitors for treating neurodegenerative diseases. Bioorg Med Chem Lett24:204-8 (2013) [PubMed] Article