| Reaction Details |
|---|
 | Report a problem with these data |
| Target | cAMP-specific 3',5'-cyclic phosphodiesterase 4A |
|---|
| Ligand | BDBM50067186 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1463981 (CHEMBL3405815) |
|---|
| IC50 | 3700±n/a nM |
|---|
| Citation |  Sawant, SD; Lakshma Reddy, G; Dar, MI; Srinivas, M; Gupta, G; Sahu, PK; Mahajan, P; Nargotra, A; Singh, S; Sharma, SC; Tikoo, M; Singh, G; Vishwakarma, RA; Syed, SH Discovery of novel pyrazolopyrimidinone analogs as potent inhibitors of phosphodiesterase type-5. Bioorg Med Chem23:2121-8 (2015) [PubMed] Article Sawant, SD; Lakshma Reddy, G; Dar, MI; Srinivas, M; Gupta, G; Sahu, PK; Mahajan, P; Nargotra, A; Singh, S; Sharma, SC; Tikoo, M; Singh, G; Vishwakarma, RA; Syed, SH Discovery of novel pyrazolopyrimidinone analogs as potent inhibitors of phosphodiesterase type-5. Bioorg Med Chem23:2121-8 (2015) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| cAMP-specific 3',5'-cyclic phosphodiesterase 4A |
|---|
| Name: | cAMP-specific 3',5'-cyclic phosphodiesterase 4A |
|---|
| Synonyms: | 3',5'-cyclic phosphodiesterase | DPDE2 | PDE46 | PDE4A | PDE4A_HUMAN | Phosphodiesterase 4 (PDE4) | Phosphodiesterase 4A | Phosphodiesterase 4A (PDE4) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 98113.27 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P27815 |
|---|
| Residue: | 886 |
|---|
| Sequence: | MEPPTVPSERSLSLSLPGPREGQATLKPPPQHLWRQPRTPIRIQQRGYSDSAERAERERQ
PHRPIERADAMDTSDRPGLRTTRMSWPSSFHGTGTGSGGAGGGSSRRFEAENGPTPSPGR
SPLDSQASPGLVLHAGAATSQRRESFLYRSDSDYDMSPKTMSRNSSVTSEAHAEDLIVTP
FAQVLASLRSVRSNFSLLTNVPVPSNKRSPLGGPTPVCKATLSEETCQQLARETLEELDW
CLEQLETMQTYRSVSEMASHKFKRMLNRELTHLSEMSRSGNQVSEYISTTFLDKQNEVEI
PSPTMKEREKQQAPRPRPSQPPPPPVPHLQPMSQITGLKKLMHSNSLNNSNIPRFGVKTD
QEELLAQELENLNKWGLNIFCVSDYAGGRSLTCIMYMIFQERDLLKKFRIPVDTMVTYML
TLEDHYHADVAYHNSLHAADVLQSTHVLLATPALDAVFTDLEILAALFAAAIHDVDHPGV
SNQFLINTNSELALMYNDESVLENHHLAVGFKLLQEDNCDIFQNLSKRQRQSLRKMVIDM
VLATDMSKHMTLLADLKTMVETKKVTSSGVLLLDNYSDRIQVLRNMVHCADLSNPTKPLE
LYRQWTDRIMAEFFQQGDRERERGMEISPMCDKHTASVEKSQVGFIDYIVHPLWETWADL
VHPDAQEILDTLEDNRDWYYSAIRQSPSPPPEEESRGPGHPPLPDKFQFELTLEEEEEEE
ISMAQIPCTAQEALTAQGLSGVEEALDATIAWEASPAQESLEVMAQEASLEAELEAVYLT
QQAQSTGSAPVAPDEFSSREEFVVAVSHSSPSALALQSPLLPAWRTLSVSEHAPGLPGLP
STAAEVEAQREHQAAKRACSACAGTFGEDTSALPAPGGGGSGGDPT
|
|
|
|---|
| BDBM50067186 |
|---|
| n/a |
|---|
| Name | BDBM50067186 |
|---|
| Synonyms: | CHEMBL3401745 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H33N5O5S |
|---|
| Mol. Mass. | 491.604 |
|---|
| SMILES | CCCc1nn(C)c2c1nc([nH]c2=O)-c1cc(ccc1OCC)S(=O)(=O)NCCCCCCO |
|---|
| Structure | 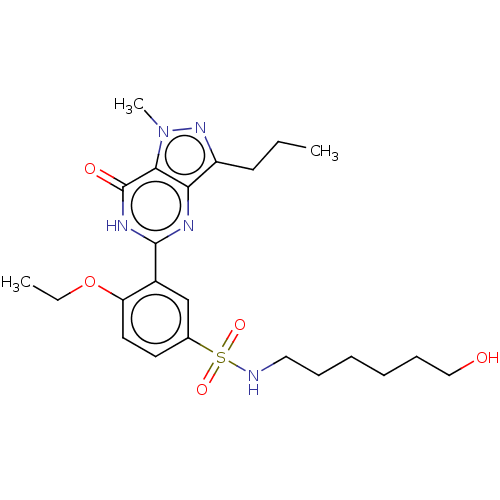
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Sawant, SD; Lakshma Reddy, G; Dar, MI; Srinivas, M; Gupta, G; Sahu, PK; Mahajan, P; Nargotra, A; Singh, S; Sharma, SC; Tikoo, M; Singh, G; Vishwakarma, RA; Syed, SH Discovery of novel pyrazolopyrimidinone analogs as potent inhibitors of phosphodiesterase type-5. Bioorg Med Chem23:2121-8 (2015) [PubMed] Article
Sawant, SD; Lakshma Reddy, G; Dar, MI; Srinivas, M; Gupta, G; Sahu, PK; Mahajan, P; Nargotra, A; Singh, S; Sharma, SC; Tikoo, M; Singh, G; Vishwakarma, RA; Syed, SH Discovery of novel pyrazolopyrimidinone analogs as potent inhibitors of phosphodiesterase type-5. Bioorg Med Chem23:2121-8 (2015) [PubMed] Article