

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Interleukin-1 receptor-associated kinase 4 | ||
| Ligand | BDBM50067533 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_1465696 (CHEMBL3405457) | ||
| Ki | 2700±n/a nM | ||
| Citation |  Boyd, MJ; Bandarage, UK; Bennett, H; Byrn, RR; Davies, I; Gu, W; Jacobs, M; Ledeboer, MW; Ledford, B; Leeman, JR; Perola, E; Wang, T; Bennani, Y; Clark, MP; Charifson, PS Isosteric replacements of the carboxylic acid of drug candidate VX-787: Effect of charge on antiviral potency and kinase activity of azaindole-based influenza PB2 inhibitors. Bioorg Med Chem Lett25:1990-4 (2015) [PubMed] Article Boyd, MJ; Bandarage, UK; Bennett, H; Byrn, RR; Davies, I; Gu, W; Jacobs, M; Ledeboer, MW; Ledford, B; Leeman, JR; Perola, E; Wang, T; Bennani, Y; Clark, MP; Charifson, PS Isosteric replacements of the carboxylic acid of drug candidate VX-787: Effect of charge on antiviral potency and kinase activity of azaindole-based influenza PB2 inhibitors. Bioorg Med Chem Lett25:1990-4 (2015) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Interleukin-1 receptor-associated kinase 4 | |||
| Name: | Interleukin-1 receptor-associated kinase 4 | ||
| Synonyms: | IRAK-4 | IRAK4 | IRAK4_HUMAN | Interleukin-1 receptor-associated kinase 4 (IRAK-4) | Interleukin-1 receptor-associated kinase 4 (IRAK4) | Renal carcinoma antigen NY-REN-64 | ||
| Type: | Protein | ||
| Mol. Mass.: | 51519.08 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Q9NWZ3 | ||
| Residue: | 460 | ||
| Sequence: |
| ||
| BDBM50067533 | |||
| n/a | |||
| Name | BDBM50067533 | ||
| Synonyms: | CHEMBL3401984 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C21H23F2N5O2 | ||
| Mol. Mass. | 415.4364 | ||
| SMILES | [H][C@]1(C(O)CO)C2CCC(CC2)[C@@H]1Nc1nc(ncc1F)-c1c[nH]c2ncc(F)cc12 |r,wD:12.14,1.0,TLB:2:1:8.7:10.11,THB:13:12:8.7:10.11,(-.64,1.55,;.13,.87,;1.58,1.51,;2.6,.82,;1.69,3.04,;2.79,3.59,;1.39,,;2.86,.48,;2.23,-.94,;,-.67,;-.29,-2.25,;1.15,-1.65,;-1.3,.28,;-2.82,.5,;-3.77,-.71,;-3.2,-2.14,;-4.15,-3.35,;-5.68,-3.13,;-6.25,-1.7,;-5.3,-.49,;-5.76,.65,;-3.58,-4.78,;-4.4,-6.04,;-3.43,-7.23,;-2,-6.66,;-.62,-7.36,;.67,-6.49,;.57,-4.96,;1.59,-4.28,;-.83,-4.26,;-2.1,-5.13,)| | ||
| Structure | 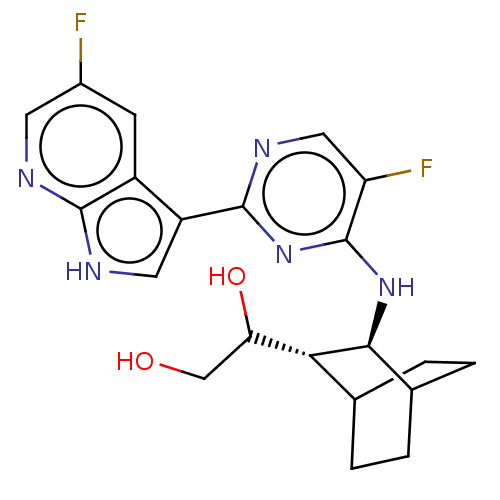
| ||