null
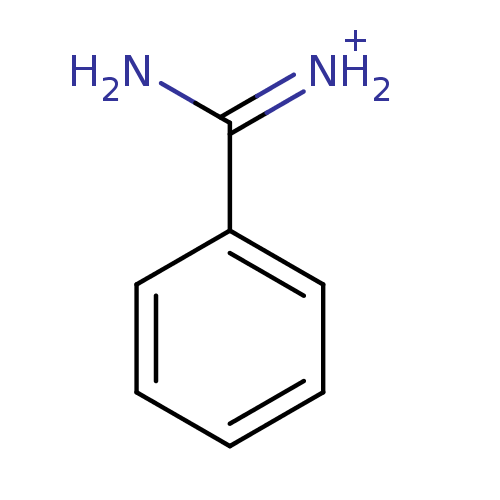
SMILES NC(=[NH2+])c1ccccc1
InChI Key InChIKey=PXXJHWLDUBFPOL-UHFFFAOYSA-O
PDB links: 40 PDB IDs match this monomer. 103 PDB IDs contain this monomer as substructures. 103 PDB IDs contain inhibitors having a similarity of 90% to this monomer.
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 17 hits for monomerid = 772
Found 17 hits for monomerid = 772
Affinity DataKi: 4.80E+3nM ΔG°: -7.18kcal/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
In DepthDetails
Article
BindingDB Entry DOI: 10.7270/Q2FX77PGPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)
BindingDB Entry DOI: 10.7270/Q2FX77PGPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)Affinity DataKi: 5.00E+3nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nM ΔG°: -6.34kcal/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nM ΔG°: -6.31kcal/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
In DepthDetails
Article
BindingDB Entry DOI: 10.7270/Q2NG4NWCPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)
BindingDB Entry DOI: 10.7270/Q2NG4NWCPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)Affinity DataKi: 2.90E+4nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 3.90E+4nM ΔG°: -6.01kcal/molepH: 8.0 T: 2°CAssay Description:The measurements were carried out on a microplate reader. Two concentrations of the substrate and five concentrations of the inhibitor were used in t...More data for this Ligand-Target Pair
In DepthDetails
Article
BindingDB Entry DOI: 10.7270/Q2445JRMPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)
BindingDB Entry DOI: 10.7270/Q2445JRMPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)Affinity DataKi: 9.50E+4nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 9.70E+4nM ΔG°: -5.42kcal/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+5nM ΔG°: -5.34kcal/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.65E+5nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+5nM ΔG°: -5.11kcal/molepH: 8.0 T: 2°CAssay Description:The measurements were carried out on a microplate reader. Two concentrations of the substrate and five concentrations of the inhibitor were used in t...More data for this Ligand-Target Pair
Affinity DataKi: 3.20E+5nM ΔG°: -4.72kcal/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 3.80E+5nMAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 6.03E+5nM ΔG°: -4.40kcal/molepH: 8.05 T: 2°CAssay Description:All readings were made in a Hitachi-Coleman ultraviolet-visible 101 spectrophotometer.More data for this Ligand-Target Pair
Affinity DataKi: 7.50E+5nM ΔG°: -4.22kcal/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+6nM ΔG°: >-4.09kcal/molepH: 8.0 T: 2°CAssay Description:The measurements were carried out on a microplate reader. Two concentrations of the substrate and five concentrations of the inhibitor were used in t...More data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+5nMpH: 8.0 T: 2°CAssay Description:The cleavage of the substrate was followed by monitoring the change in fluorescence at 460 nm (excitation at 365 nm) for 25 min at room temperature o...More data for this Ligand-Target Pair
Activity Spreadsheet -- ITC Data from BindingDB
 Found 4 hits for monomerid = 772
Found 4 hits for monomerid = 772
ITC DataΔG°: -6.42kcal/mole −TΔS°: -2.37kcal/mole ΔH°: -4.06kcal/mole logk: 6.20E+4
pH: 8.0 T: 20.10°C
pH: 8.0 T: 20.10°C
ITC DataΔG°: -6.38kcal/mole −TΔS°: -1.29kcal/mole ΔH°: -5.11kcal/mole logk: 4.10E+4
pH: 8.0 T: 30.00°C
pH: 8.0 T: 30.00°C
ITC DataΔG°: -6.38kcal/mole −TΔS°: -0.719kcal/mole ΔH°: -5.66kcal/mole logk: 3.10E+4
pH: 8.0 T: 37.10°C
pH: 8.0 T: 37.10°C
ITC DataΔG°: -6.35kcal/mole −TΔS°: -1.84kcal/mole ΔH°: -4.51kcal/mole logk: 4.50E+4
pH: 8.0 T: 25.10°C
pH: 8.0 T: 25.10°C