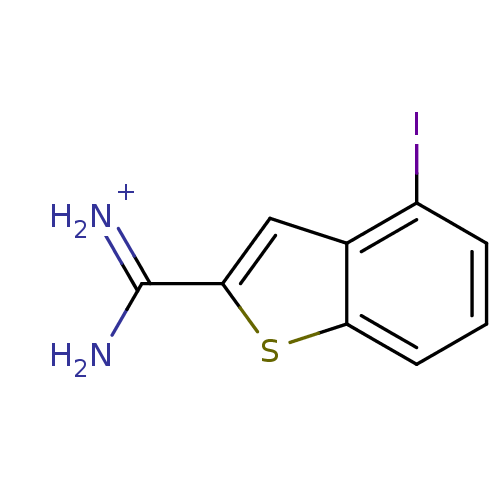
Affinity DataKi: 210nM ΔG°: -37.7kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
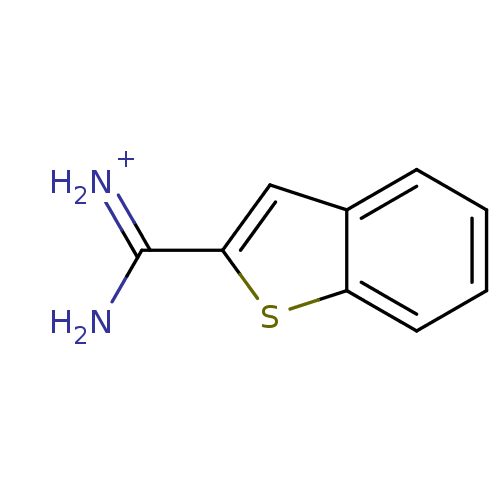
Affinity DataKi: 440nM ΔG°: -35.9kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
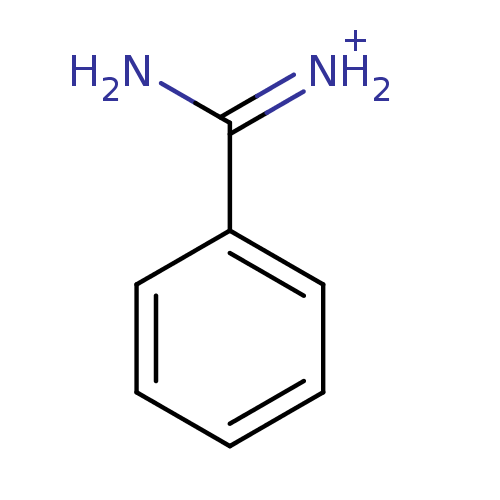
Affinity DataKi: 1.00E+3nM ΔG°: -33.9kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
In DepthDetails
Article
BindingDB Entry DOI: 10.7270/Q2NG4NWCPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)
BindingDB Entry DOI: 10.7270/Q2NG4NWCPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)Affinity DataKi: 1.50E+3nM ΔG°: -32.9kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nM ΔG°: -31.9kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 9.90E+3nM ΔG°: -28.3kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+4nM ΔG°: -27.3kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.68E+4nM ΔG°: -27.0kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nM ΔG°: -26.6kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nM ΔG°: -26.6kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nM ΔG°: -26.4kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nM ΔG°: -26.4kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
In DepthDetails
Article
BindingDB Entry DOI: 10.7270/Q2NG4NWCPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)
BindingDB Entry DOI: 10.7270/Q2NG4NWCPubMedDrugBank
MMDB
PDB
 3D Structure (crystal)
3D Structure (crystal)Affinity DataKi: 3.00E+4nM ΔG°: -25.6kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 3.40E+4nM ΔG°: -25.2kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 5.80E+4nM ΔG°: -23.9kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 6.30E+4nM ΔG°: -23.7kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 8.00E+4nM ΔG°: -23.1kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 9.70E+4nM ΔG°: -22.7kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+5nM ΔG°: -22.4kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 3.20E+5nM ΔG°: -19.7kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 3.60E+5nM ΔG°: -19.5kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 7.50E+5nM ΔG°: -17.7kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+6nM ΔG°: -15.8kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+6nM ΔG°: -15.5kJ/molepH: 7.4 T: 2°CAssay Description:Each enzyme was assayed with a set of different concentrations of each inhibitor. After addition of the appropriate substrate, the rate of hydrolysis...More data for this Ligand-Target Pair