

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kJ/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Adenosine deaminase (Homo sapiens (Human)) | BDBM22920 (1-[(2R)-1-hydroxy-4-{6-[3-(1-methyl-1H-1,3-benzodi...) | PDB MMDB NCI pathway Reactome pathway KEGG UniProtKB/SwissProt B.MOAD DrugBank antibodypedia GoogleScholar AffyNet | CHEMBL MMDB PC cid PC sid PDB UniChem Similars | MMDB PDB Article PubMed | 7.70 | -45.8 | n/a | n/a | n/a | n/a | n/a | 7.4 | 22 |
Fujisawa Pharmaceutical Co., Ltd. | Assay Description The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a... | J Med Chem 47: 2728-31 (2004) Article DOI: 10.1021/jm0499559 BindingDB Entry DOI: 10.7270/Q22F7KQT | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| Adenosine deaminase (Homo sapiens (Human)) | BDBM22950 (1-[(3R,4S)-4-hydroxy-1-(naphthalen-2-yloxy)pentan-...) | PDB MMDB NCI pathway Reactome pathway KEGG UniProtKB/SwissProt B.MOAD DrugBank antibodypedia GoogleScholar AffyNet | DrugBank MMDB PC cid PC sid PDB UniChem | MMDB PDB Article PubMed | 9.80 | -45.3 | n/a | n/a | n/a | n/a | n/a | 7.4 | 22 |
Fujisawa Pharmaceutical Co., Ltd. | Assay Description The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a... | J Med Chem 47: 2728-31 (2004) Article DOI: 10.1021/jm0499559 BindingDB Entry DOI: 10.7270/Q22F7KQT | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| Adenosine deaminase (Homo sapiens (Human)) | BDBM22948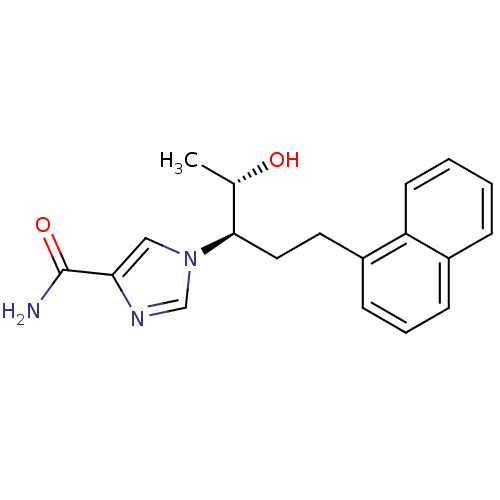 (1-[(1R,2S)-2-hydroxy-1-(2-naphthalen-1-ylethyl)pro...) | PDB MMDB NCI pathway Reactome pathway KEGG UniProtKB/SwissProt B.MOAD DrugBank antibodypedia GoogleScholar AffyNet | Purchase CHEMBL DrugBank MCE MMDB PC cid PC sid PDB UniChem Similars | MMDB PDB Article PubMed | 11 | -45.0 | n/a | n/a | n/a | n/a | n/a | 7.4 | 22 |
Fujisawa Pharmaceutical Co., Ltd. | Assay Description The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a... | J Med Chem 47: 2728-31 (2004) Article DOI: 10.1021/jm0499559 BindingDB Entry DOI: 10.7270/Q22F7KQT | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| Adenosine deaminase (Homo sapiens (Human)) | BDBM22949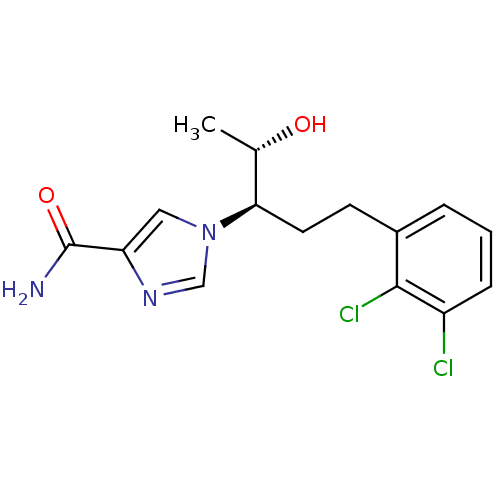 (1-[(3R,4S)-1-(2,3-dichlorophenyl)-4-hydroxypentan-...) | PDB MMDB NCI pathway Reactome pathway KEGG UniProtKB/SwissProt B.MOAD DrugBank antibodypedia GoogleScholar AffyNet | CHEMBL DrugBank MMDB PC cid PC sid PDB UniChem Patents | MMDB PDB Article PubMed | 13 | -44.6 | n/a | n/a | n/a | n/a | n/a | 7.4 | 22 |
Fujisawa Pharmaceutical Co., Ltd. | Assay Description The reaction velocity was measured by change in absorbance at 265nm (A265) resulting from the deamination of adenosine. The reaction was started by a... | J Med Chem 47: 2728-31 (2004) Article DOI: 10.1021/jm0499559 BindingDB Entry DOI: 10.7270/Q22F7KQT | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||