| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Retinoic acid receptor RXR-alpha |
|---|
| Ligand | BDBM50218451 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_440654 (CHEMBL889748) |
|---|
| IC50 | 720±n/a nM |
|---|
| Citation |  Sakaki, J; Kishida, M; Konishi, K; Gunji, H; Toyao, A; Matsumoto, Y; Kanazawa, T; Uchiyama, H; Fukaya, H; Mitani, H; Arai, Y; Kimura, M Synthesis and structure-activity relationship of novel RXR antagonists: orally active anti-diabetic and anti-obesity agents. Bioorg Med Chem Lett17:4804-7 (2007) [PubMed] Article Sakaki, J; Kishida, M; Konishi, K; Gunji, H; Toyao, A; Matsumoto, Y; Kanazawa, T; Uchiyama, H; Fukaya, H; Mitani, H; Arai, Y; Kimura, M Synthesis and structure-activity relationship of novel RXR antagonists: orally active anti-diabetic and anti-obesity agents. Bioorg Med Chem Lett17:4804-7 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Retinoic acid receptor RXR-alpha |
|---|
| Name: | Retinoic acid receptor RXR-alpha |
|---|
| Synonyms: | NR2B1 | Nuclear receptor subfamily 2 group B member 1 | Nuclear receptor subfamily 4 group A member 2 | Nuclear receptor subfamily 4 group A member 2/Retinoic acid receptor RXR-alpha | RXRA | RXRA_HUMAN | Retinoic acid receptor RXR-alpha/gamma | Retinoid X receptor alpha | Retinoid receptor |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 50820.38 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ChEMBL_1456363 |
|---|
| Residue: | 462 |
|---|
| Sequence: | MDTKHFLPLDFSTQVNSSLTSPTGRGSMAAPSLHPSLGPGIGSPGQLHSPISTLSSPING
MGPPFSVISSPMGPHSMSVPTTPTLGFSTGSPQLSSPMNPVSSSEDIKPPLGLNGVLKVP
AHPSGNMASFTKHICAICGDRSSGKHYGVYSCEGCKGFFKRTVRKDLTYTCRDNKDCLID
KRQRNRCQYCRYQKCLAMGMKREAVQEERQRGKDRNENEVESTSSANEDMPVERILEAEL
AVEPKTETYVEANMGLNPSSPNDPVTNICQAADKQLFTLVEWAKRIPHFSELPLDDQVIL
LRAGWNELLIASFSHRSIAVKDGILLATGLHVHRNSAHSAGVGAIFDRVLTELVSKMRDM
QMDKTELGCLRAIVLFNPDSKGLSNPAEVEALREKVYASLEAYCKHKYPEQPGRFAKLLL
RLPALRSIGLKCLEHLFFFKLIGDTPIDTFLMEMLEAPHQMT
|
|
|
|---|
| BDBM50218451 |
|---|
| n/a |
|---|
| Name | BDBM50218451 |
|---|
| Synonyms: | 4-[2-(4-methoxy-phenylsulfanyl)-5,7,7,10,10-pentamethyl-7,8,9,10-tetrahydro-5H-5,13-diaza-benzo[4,5]cyclohepta[1,2-b]naphthalen-12-yl]-benzoic acid | CHEMBL395563 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C36H36N2O3S |
|---|
| Mol. Mass. | 576.748 |
|---|
| SMILES | COc1ccc(Sc2ccc3N(C)c4cc5c(cc4C(=Nc3c2)c2ccc(cc2)C(O)=O)C(C)(C)CCC5(C)C)cc1 |c:20| |
|---|
| Structure | 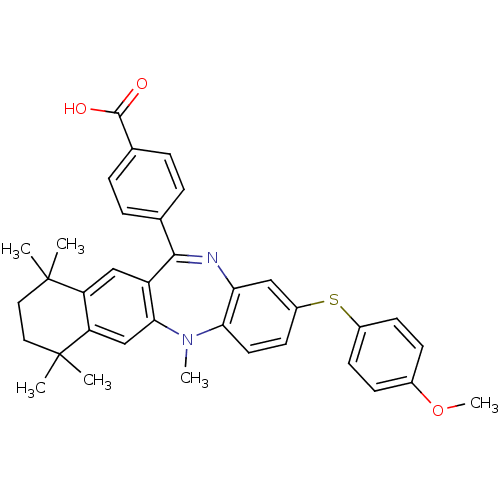
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Sakaki, J; Kishida, M; Konishi, K; Gunji, H; Toyao, A; Matsumoto, Y; Kanazawa, T; Uchiyama, H; Fukaya, H; Mitani, H; Arai, Y; Kimura, M Synthesis and structure-activity relationship of novel RXR antagonists: orally active anti-diabetic and anti-obesity agents. Bioorg Med Chem Lett17:4804-7 (2007) [PubMed] Article
Sakaki, J; Kishida, M; Konishi, K; Gunji, H; Toyao, A; Matsumoto, Y; Kanazawa, T; Uchiyama, H; Fukaya, H; Mitani, H; Arai, Y; Kimura, M Synthesis and structure-activity relationship of novel RXR antagonists: orally active anti-diabetic and anti-obesity agents. Bioorg Med Chem Lett17:4804-7 (2007) [PubMed] Article