| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Lysine-specific demethylase 6B |
|---|
| Ligand | BDBM50513344 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1853070 (CHEMBL4353694) |
|---|
| IC50 | >100000±n/a nM |
|---|
| Citation |  Le Bihan, YV; Lanigan, RM; Atrash, B; McLaughlin, MG; Velupillai, S; Malcolm, AG; England, KS; Ruda, GF; Mok, NY; Tumber, A; Tomlin, K; Saville, H; Shehu, E; McAndrew, C; Carmichael, L; Bennett, JM; Jeganathan, F; Eve, P; Donovan, A; Hayes, A; Wood, F; Raynaud, FI; Fedorov, O; Brennan, PE; Burke, R; van Montfort, RLM; Rossanese, OW; Blagg, J; Bavetsias, V C8-substituted pyrido[3,4-d]pyrimidin-4(3H)-ones: Studies towards the identification of potent, cell penetrant Jumonji C domain containing histone lysine demethylase 4 subfamily (KDM4) inhibitors, compound profiling in cell-based target engagement assays. Eur J Med Chem177:316-337 (2019) [PubMed] Article Le Bihan, YV; Lanigan, RM; Atrash, B; McLaughlin, MG; Velupillai, S; Malcolm, AG; England, KS; Ruda, GF; Mok, NY; Tumber, A; Tomlin, K; Saville, H; Shehu, E; McAndrew, C; Carmichael, L; Bennett, JM; Jeganathan, F; Eve, P; Donovan, A; Hayes, A; Wood, F; Raynaud, FI; Fedorov, O; Brennan, PE; Burke, R; van Montfort, RLM; Rossanese, OW; Blagg, J; Bavetsias, V C8-substituted pyrido[3,4-d]pyrimidin-4(3H)-ones: Studies towards the identification of potent, cell penetrant Jumonji C domain containing histone lysine demethylase 4 subfamily (KDM4) inhibitors, compound profiling in cell-based target engagement assays. Eur J Med Chem177:316-337 (2019) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Lysine-specific demethylase 6B |
|---|
| Name: | Lysine-specific demethylase 6B |
|---|
| Synonyms: | JMJD3 | JmjC domain-containing protein 3 | Jumonji domain-containing protein 3 | KDM6B | KDM6B_HUMAN | KIAA0346 | Lysine demethylase 6B | Lysine-specific demethylase 6B | Lysine-specific demethylase 6B (KDM6B(JMJD3)) | Lysine-specific demethylase 6B (KDM6B) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 176672.55 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | O15054 |
|---|
| Residue: | 1643 |
|---|
| Sequence: | MHRAVDPPGARAAREAFALGGLSCAGAWSSCPPHPPPRSAWLPGGRCSASIGQPPLPAPL
PPSHGSSSGHPSKPYYAPGAPTPRPLHGKLESLHGCVQALLREPAQPGLWEQLGQLYESE
HDSEEATRCYHSALRYGGSFAELGPRIGRLQQAQLWNFHTGSCQHRAKVLPPLEQVWNLL
HLEHKRNYGAKRGGPPVKRAAEPPVVQPVPPAALSGPSGEEGLSPGGKRRRGCNSEQTGL
PPGLPLPPPPLPPPPPPPPPPPPPLPGLATSPPFQLTKPGLWSTLHGDAWGPERKGSAPP
ERQEQRHSLPHPYPYPAPAYTAHPPGHRLVPAAPPGPGPRPPGAESHGCLPATRPPGSDL
RESRVQRSRMDSSVSPAATTACVPYAPSRPPGLPGTTTSSSSSSSSNTGLRGVEPNPGIP
GADHYQTPALEVSHHGRLGPSAHSSRKPFLGAPAATPHLSLPPGPSSPPPPPCPRLLRPP
PPPAWLKGPACRAAREDGEILEELFFGTEGPPRPAPPPLPHREGFLGPPASRFSVGTQDS
HTPPTPPTPTTSSSNSNSGSHSSSPAGPVSFPPPPYLARSIDPLPRPPSPAQNPQDPPLV
PLTLALPPAPPSSCHQNTSGSFRRPESPRPRVSFPKTPEVGPGPPPGPLSKAPQPVPPGV
GELPARGPRLFDFPPTPLEDQFEEPAEFKILPDGLANIMKMLDESIRKEEEQQQHEAGVA
PQPPLKEPFASLQSPFPTDTAPTTTAPAVAVTTTTTTTTTTTATQEEEKKPPPALPPPPP
LAKFPPPSQPQPPPPPPPSPASLLKSLASVLEGQKYCYRGTGAAVSTRPGPLPTTQYSPG
PPSGATALPPTSAAPSAQGSPQPSASSSSQFSTSGGPWARERRAGEEPVPGPMTPTQPPP
PLSLPPARSESEVLEEISRACETLVERVGRSATDPADPVDTAEPADSGTERLLPPAQAKE
EAGGVAAVSGSCKRRQKEHQKEHRRHRRACKDSVGRRPREGRAKAKAKVPKEKSRRVLGN
LDLQSEEIQGREKSRPDLGGASKAKPPTAPAPPSAPAPSAQPTPPSASVPGKKAREEAPG
PPGVSRADMLKLRSLSEGPPKELKIRLIKVESGDKETFIASEVEERRLRMADLTISHCAA
DVVRASRNAKVKGKFRESYLSPAQSVKPKINTEEKLPREKLNPPTPSIYLESKRDAFSPV
LLQFCTDPRNPITVIRGLAGSLRLNLGLFSTKTLVEASGEHTVEVRTQVQQPSDENWDLT
GTRQIWPCESSRSHTTIAKYAQYQASSFQESLQEEKESEDEESEEPDSTTGTPPSSAPDP
KNHHIIKFGTNIDLSDAKRWKPQLQELLKLPAFMRVTSTGNMLSHVGHTILGMNTVQLYM
KVPGSRTPGHQENNNFCSVNINIGPGDCEWFAVHEHYWETISAFCDRHGVDYLTGSWWPI
LDDLYASNIPVYRFVQRPGDLVWINAGTVHWVQATGWCNNIAWNVGPLTAYQYQLALERY
EWNEVKNVKSIVPMIHVSWNVARTVKISDPDLFKMIKFCLLQSMKHCQVQRESLVRAGKK
IAYQGRVKDEPAYYCNECDVEVFNILFVTSENGSRNTYLVHCEGCARRRSAGLQGVVVLE
QYRTEELAQAYDAFTLAPASTSR
|
|
|
|---|
| BDBM50513344 |
|---|
| n/a |
|---|
| Name | BDBM50513344 |
|---|
| Synonyms: | CHEMBL4447515 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H23FN6O |
|---|
| Mol. Mass. | 418.4667 |
|---|
| SMILES | Fc1cccc(c1)C1CCN(CCc2cnn(c2)-c2nccc3c2nc[nH]c3=O)CC1 |
|---|
| Structure | 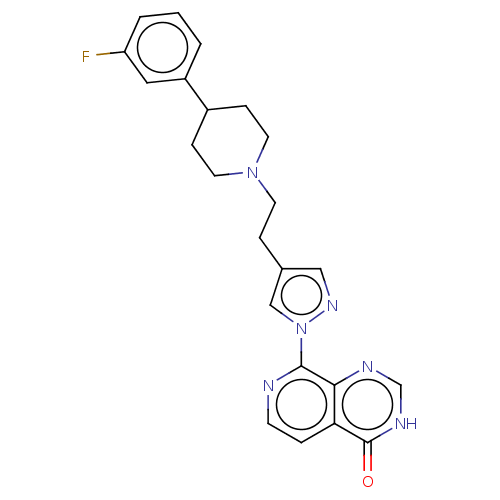
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Le Bihan, YV; Lanigan, RM; Atrash, B; McLaughlin, MG; Velupillai, S; Malcolm, AG; England, KS; Ruda, GF; Mok, NY; Tumber, A; Tomlin, K; Saville, H; Shehu, E; McAndrew, C; Carmichael, L; Bennett, JM; Jeganathan, F; Eve, P; Donovan, A; Hayes, A; Wood, F; Raynaud, FI; Fedorov, O; Brennan, PE; Burke, R; van Montfort, RLM; Rossanese, OW; Blagg, J; Bavetsias, V C8-substituted pyrido[3,4-d]pyrimidin-4(3H)-ones: Studies towards the identification of potent, cell penetrant Jumonji C domain containing histone lysine demethylase 4 subfamily (KDM4) inhibitors, compound profiling in cell-based target engagement assays. Eur J Med Chem177:316-337 (2019) [PubMed] Article
Le Bihan, YV; Lanigan, RM; Atrash, B; McLaughlin, MG; Velupillai, S; Malcolm, AG; England, KS; Ruda, GF; Mok, NY; Tumber, A; Tomlin, K; Saville, H; Shehu, E; McAndrew, C; Carmichael, L; Bennett, JM; Jeganathan, F; Eve, P; Donovan, A; Hayes, A; Wood, F; Raynaud, FI; Fedorov, O; Brennan, PE; Burke, R; van Montfort, RLM; Rossanese, OW; Blagg, J; Bavetsias, V C8-substituted pyrido[3,4-d]pyrimidin-4(3H)-ones: Studies towards the identification of potent, cell penetrant Jumonji C domain containing histone lysine demethylase 4 subfamily (KDM4) inhibitors, compound profiling in cell-based target engagement assays. Eur J Med Chem177:316-337 (2019) [PubMed] Article