| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Thymidine phosphorylase |
|---|
| Ligand | BDBM20079 |
|---|
| Substrate/Competitor | BDBM1 |
|---|
| Meas. Tech. | Enzyme Inhibition Assay |
|---|
| pH | 6.4±n/a |
|---|
| Temperature | 310.15±n/a K |
|---|
| Ki | 1.3±0.17 nM |
|---|
| Km | 121000±19000 nM |
|---|
| Comments | Data was taken from Biochem. Pharmacol. 2000, 59, 1227-1236. |
|---|
| Citation |  Nencka, R; Votruba, I; Hrebabecký, H; Jansa, P; Tloust'ova, E; Horska, K; Masojídkova, M; Holý, A Discovery of 5-Substituted-6-chlorouracils as Efficient Inhibitors of Human Thymidine Phosphorylase. J Med Chem50:6016-23 (2007) [PubMed] Article Nencka, R; Votruba, I; Hrebabecký, H; Jansa, P; Tloust'ova, E; Horska, K; Masojídkova, M; Holý, A Discovery of 5-Substituted-6-chlorouracils as Efficient Inhibitors of Human Thymidine Phosphorylase. J Med Chem50:6016-23 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Thymidine phosphorylase |
|---|
| Name: | Thymidine phosphorylase |
|---|
| Synonyms: | ECGF1 | Gliostatin | PD-ECGF | Platelet-derived endothelial cell growth factor | TP | TYMP | TYPH_HUMAN | TdRPase | Thymidine Phosphorylase (TP) | Thymidine phosphorylase |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 49948.87 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | The recombinant human thymidine phosphorylase (V79TP) was expressed in V79 Chinese hamster cells (Sigma, T-9319). |
|---|
| Residue: | 482 |
|---|
| Sequence: | MAALMTPGTGAPPAPGDFSGEGSQGLPDPSPEPKQLPELIRMKRDGGRLSEADIRGFVAA
VVNGSAQGAQIGAMLMAIRLRGMDLEETSVLTQALAQSGQQLEWPEAWRQQLVDKHSTGG
VGDKVSLVLAPALAACGCKVPMISGRGLGHTGGTLDKLESIPGFNVIQSPEQMQVLLDQA
GCCIVGQSEQLVPADGILYAARDVTATVDSLPLITASILSKKLVEGLSALVVDVKFGGAA
VFPNQEQARELAKTLVGVGASLGLRVAAALTAMDKPLGRCVGHALEVEEALLCMDGAGPP
DLRDLVTTLGGALLWLSGHAGTQAQGAARVAAALDDGSALGRFERMLAAQGVDPGLARAL
CSGSPAERRQLLPRAREQEELLAPADGTVELVRALPLALVLHELGAGRSRAGEPLRLGVG
AELLVDVGQRLRRGTPWLRVHRDGPALSGPQSRALQEALVLSDRAPFAAPSPFAELVLPP
QQ
|
|
|
|---|
| BDBM20079 |
|---|
| BDBM1 |
|---|
| Name | BDBM20079 |
|---|
| Synonyms: | 5-chloro-6-[(2-iminopyrrolidin-1-yl)methyl]-1,2,3,4-tetrahydropyrimidine-2,4-dione | 5-chloro-6-[(2-iminopyrrolidin-1-yl)methyl]pyrimidine-2,4(1H,3H)-dione | CHEMBL65375 | Lonsurf | TPI | Tipiracil |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C9H11ClN4O2 |
|---|
| Mol. Mass. | 242.662 |
|---|
| SMILES | Clc1c(CN2CCCC2=N)[nH]c(=O)[nH]c1=O |
|---|
| Structure | 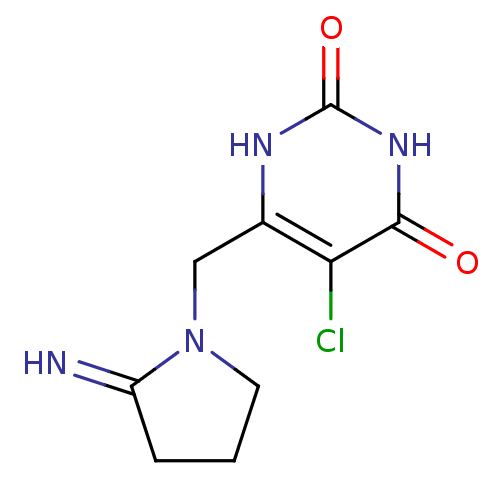
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Nencka, R; Votruba, I; Hrebabecký, H; Jansa, P; Tloust'ova, E; Horska, K; Masojídkova, M; Holý, A Discovery of 5-Substituted-6-chlorouracils as Efficient Inhibitors of Human Thymidine Phosphorylase. J Med Chem50:6016-23 (2007) [PubMed] Article
Nencka, R; Votruba, I; Hrebabecký, H; Jansa, P; Tloust'ova, E; Horska, K; Masojídkova, M; Holý, A Discovery of 5-Substituted-6-chlorouracils as Efficient Inhibitors of Human Thymidine Phosphorylase. J Med Chem50:6016-23 (2007) [PubMed] Article